News
Newswise — Exercise is powerful medicine for physical and mental wellbeing. But people’s ability to harness proven benefits from exercise have been greatly challenged during this pandemic, according to Thomas M. Best, M.D., Ph.D., FACSM, professor of orthopedic surgery at the University of Miami Miller School of Medicine and research director of the UHealth Sports Medicine Institute.
“This pandemic has limited, curtailed and dramatically altered people’s ability to sustain physical activity. Many haven’t exercised for months due to lockdowns, weather, safety issues, illness, and more,” said Dr. Best, former president of the American College of Sports Medicine, a former Olympic athlete and today’s team physician for the University of Miami Hurricanes and Miami Marlins professional baseball team.
To address and overcome the challenges so Americans can return to or sustain physical activity safely, Dr. Best and sports medicine colleagues from around the U.S. wrote “COVID-19: Considerations for Sports and Physical Activity,” published August 7 in Current Sports Medicine Reports, an American College of Sports Medicine journal.
There are multiple guidelines on sports in the era of COVID-19 but they focus on elite athletes, including college and professional athletes, according to Dr. Best.
“Until this paper, there hasn’t been anything out there for everyday people, including those who exercise regularly and those who don’t but should,” he said.
The authors point to scientific evidence showing that too little or too much exercise can impact physical and mental health. For example, there are diseases of inactivity, including obesity and hypertension. Exercising moderately for 30 to 60 minutes most days of the week can both prevent and mitigate the effects of chronic disease and improve physical and mental health, as well as enhance immune function. Overtraining, on the other hand, can have short term detrimental effects on the immune system and therefore diminish one’s ability to fight off the common cold and other illnesses.
At the right amount for an individual, physical activity is an incredibly powerful medicine to fight COVID-19 and other diseases, according to Dr. Best.
“Studies continue to mount showing that exercise has implications even for longevity, with exercisers living an average 3 years longer than non-exercisers,” Dr. Best said.
One of the challenges in making exercise recommendations is pinpointing a “sweet spot,” at which exercise does the most good and no harm.
“We don’t know what the precise dose is, yet and it may be different for everyone,” he said. “We know, however, that no exercise is detrimental to the immune system, moderate exercise enhances the immune system and too much exercise does the immune system more harm than good.”
Among the messages to primary care and other providers during the pandemic is to recommend exercise, but in a way that patients who might be unfit can and will sustain recommended amounts.
The authors’ calls to action include:
Encouraging people who are well to start or continue moderate physical activity, including cardiovascular and strength training, for 150 to 300 minutes a week.
“The greater concern is for those who haven’t exercised in recent months. Some of those people might have one or two chronic diseases, like hypertension, obesity, or diabetes. So, their return to physical activity and sports is a little trickier in terms of how providers should prescribe the exercise,” according to Dr. Best. “If you go to your physician and he or she says exercise 30 minutes a day 4 or 5 days a week, that doesn’t help a lot of people who are looking for more specifics. And as we come back from this pandemic and a period of physical inactivity, I think it’s even going to be more challenging.”
It is important, for example, that patients know they do not have to do the whole 30 minutes in one session. They can break up daily exercise into two 15-minute sessions or three 10-minute sessions and get about the same benefit, according to Dr. Best.
For people to exercise indoors or outdoors safely, they should wear masks around others and maintain 6 feet of distance. This is particularly important in group exercise activities to slow viral transmission.
Especially people at high risk for coronavirus exposure should refrain from exhaustive exercise and overtraining. And people who are returning from a period of inactivity, should not aim for a personal best in their first sports event, according to Dr. Best.
“We don’t want to wear down our immune systems because we know the immune system plays a key role in trying to fight off any kind of illness, including viruses,” he said. “Overtraining is defined by the individual. The goal is to optimize your immune system with optimal training, sleep, nutrition and hydration.”
Innovative strategies, like wearable technologies, can help people get back to and stay with regular physical activity.
Telemedicine works well in certain settings, and providers could consider telemedicine visits when assisting people getting back to regular physical activity. Telemedicine may be useful, in particular, for populations vulnerable to the COVID pandemic.
It is important to get people to exercise despite pandemic-related challenges, and this call to action paper starts that conversation, Dr. Best said.
“I think a lot of people became inactive during the pandemic and the longer inactivity goes on, the more we know that will have long-term detrimental effects on our health. We have to deal with this challenge, and meet it head on,” he said. “The University of Miami is uniquely positioned given the diverse populations and communities we serve to make a transformational contribution to understanding the coronavirus pandemic and efforts to mitigate its short-term and long-term effects. Our success at fighting COVID can only be enhanced with a robust effort to keep people safely exercising and understanding the critical role that physical activity plays in human health.”
Photo Credit: Getty Images
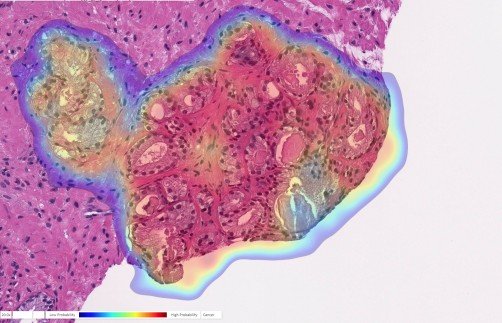
Newswise — PITTSBURGH, July 27, 2020 – A study published today in The Lancet Digital Health by UPMC and University of Pittsburgh researchers demonstrates the highest accuracy to date in recognizing and characterizing prostate cancer using an artificial intelligence (AI) program.
“Humans are good at recognizing anomalies, but they have their own biases or past experience,” said senior author Rajiv Dhir, M.D., M.B.A., chief pathologist and vice chair of pathology at UPMC Shadyside and professor of biomedical informatics at Pitt. “Machines are detached from the whole story. There’s definitely an element of standardizing care.”
To train the AI to recognize prostate cancer, Dhir and his colleagues provided images from more than a million parts of stained tissue slides taken from patient biopsies. Each image was labeled by expert pathologists to teach the AI how to discriminate between healthy and abnormal tissue. The algorithm was then tested on a separate set of 1,600 slides taken from 100 consecutive patients seen at UPMC for suspected prostate cancer.
During testing, the AI demonstrated 98% sensitivity and 97% specificity at detecting prostate cancer — significantly higher than previously reported for algorithms working from tissue slides.
Also, this is the first algorithm to extend beyond cancer detection, reporting high performance for tumor grading, sizing and invasion of the surrounding nerves. These all are clinically important features required as part of the pathology report.
AI also flagged six slides that were not noted by the expert pathologists.
But Dhir explained that this doesn’t necessarily mean that the machine is superior to humans. For example, in the course of evaluating these cases, the pathologist could have simply seen enough evidence of malignancy elsewhere in that patient’s samples to recommend treatment. For less experienced pathologists, though, the algorithm could act as a failsafe to catch cases that might otherwise be missed.
“Algorithms like this are especially useful in lesions that are atypical,” Dhir said. “A nonspecialized person may not be able to make the correct assessment. That’s a major advantage of this kind of system.”
While these results are promising, Dhir cautions that new algorithms will have to be trained to detect different types of cancer. The pathology markers aren’t universal across all tissue types. But he didn’t see why that couldn’t be done to adapt this technology to work with breast cancer, for example.
Additional authors on the study include Liron Pantanowitz, M.B.B.Ch., of the University of Michigan; Gabriela Quiroga-Garza, M.D., of UPMC; Lilach Bien, Ronen Heled, Daphna Laifenfeld, Ph.D., Chaim Linhart, Judith Sandbank, M.D., Manuela Vecsler, of Ibex Medical Analytics; Anat Albrecht-Shach, M.D., of Shamir Medical Center; Varda Shalev, M.D., M.P.A., of Maccabbi Healthcare Services; and Pamela Michelow, M.S., and Scott Hazelhurst, Ph.D., of the University of the Witwatersrand.
Funding for this study was provided by Ibex, which also created this commercially available algorithm. Pantanowitz, Shalev and Albrecht-Shach report fees paid by Ibex, and Pantanowitz and Shalev serve on the medical advisory board. Bien and Linhart are authors on pending patents US 62/743,559 and US 62/981,925. Ibex had no influence over the design of the study or the interpretation of the results.
Photo Credit:
Ibex Medical Analytics
Artificial Intelligence (AI) Detects Cancer in Prostate Biopsy: Prostate biopsy with cancer probability (blue is low, red is high). This case was originally diagnosed as benign but changed to cancer upon further review. The AI accurately detected cancer in this tricky case.
Newswise — TROY, N.Y. — In a test of antiviral effectiveness against the virus that causes COVID-19, an extract from edible seaweeds substantially outperformed remdesivir, the current standard antiviral used to combat the disease. Heparin, a common blood thinner, and a heparin variant stripped of its anticoagulant properties, performed on par with remdesivir in inhibiting SARS-CoV-2 infection in mammalian cells.
Published online today in Cell Discovery, the research is the latest example of a decoy strategy researchers from the Center for Biotechnology and Interdisciplinary Studies (CBIS) at Rensselear Polytechnic Institute are developing against viruses like the novel coronavirus that spawned the current global health crisis.
The spike protein on the surface of SARS-CoV-2 latches onto the ACE-2 receptor, a molecule on the surface of human cells. Once secured, the virus inserts its own genetic material into the cell, hijacking the cellular machinery to produce replica viruses. But the virus could just as easily be persuaded to lock onto a decoy molecule that offers a similar fit. The neutralized virus would be trapped and eventually degrade naturally.
Previous research has shown this decoy technique works in trapping other viruses, including dengue, Zika, and influenza A. To hear the researchers discuss their findings, watch this video.
“We’re learning how to block viral infection, and that is knowledge we are going to need if we want to rapidly confront pandemics,” said Jonathan Dordick, the lead researcher and a professor of chemical and biological engineering at Rensselaer Polytechnic Institute. “The reality is that we don’t have great antivirals. To protect ourselves against future pandemics, we are going to need an arsenal of approaches that we can quickly adapt to emerging viruses.”
The Cell Discovery paper tests antiviral activity in three variants of heparin (heparin, trisulfated heparin, and a non-anticoagulant low molecular weight heparin) and two fucoidans (RPI-27 and RPI-28) extracted from seaweed. All five compounds are long chains of sugar molecules known as sulfated polysaccharides, a structural conformation that the results of a binding study published earlier this month in Antiviral Research suggested as an effective decoy.
The researchers performed a dose response study known as an EC50 — shorthand for the effective concentration of the compound that inhibits 50% of viral infectivity — with each of the five compounds on mammalian cells. For the results of an EC50, which are given in a molar concentration, a lower value signals a more potent compound.
RPI-27 yielded an EC50 value of approximately 83 nanomolar, while a similar previously published and independent in vitro test of remdesivir on the same mammalian cells yielded an EC50 of 770 nanomolar. Heparin yielded an EC50 of 2.1 micromolar, or about one-third as active as remdesivir, and a non-anticoagulant analog of heparin yielded an EC50 of 5.0 micromolar, about one-fifth as active as remdesivir.
A separate test found no cellular toxicity in any of the compounds, even at the highest concentrations tested.
“What interests us is a new way of getting at infection,” said Robert Linhardt, a Rensselaer professor of chemistry and chemical biology who is collaborating with Dordick to develop the decoy strategy. “The current thinking is that the COVID-19 infection starts in the nose, and either of these substances could be the basis for a nasal spray. If you could simply treat the infection early, or even treat before you have the infection, you would have a way of blocking it before it enters the body.”
Dordick added that compounds from seaweed “could serve as a basis for an oral delivery approach to address potential gastrointestinal infection.”
In studying SARS-CoV-2 sequencing data, Dordick and Linhardt recognized several motifs on the structure of the spike protein that promised a fit compatible with heparin, a result borne out in the binding study. The spike protein is heavily encrusted in glycans, an adaptation that protects it from human enzymes which could degrade it, and prepares it to bind with a specific receptor on the cell surface.
“It’s a very complicated mechanism that we quite frankly don’t know all the details about, but we’re getting more information,” said Dordick. “One thing that’s become clear with this study is that the larger the molecule, the better the fit. The more successful compounds are the larger sulfated polysaccharides that offer a greater number of sites on the molecules to trap the virus.”
Molecular modeling based on the binding study revealed sites on the spike protein where the heparin was able to interact, raising the prospects for similar sulfated polysaccharides.
“This exciting research by Professors Dordick and Linhardt is among several ongoing research efforts at CBIS, as well as elsewhere at Rensselaer, to tackle the challenges of the COVID-19 pandemic through novel therapeutic approaches and the repurposing of existing drugs,” said CBIS Director Deepak Vashishth.
“Sulfated polysaccharides effectively inhibit SARS-CoV-2 in vitro” was published in Cell Discovery with the support of the National Research Foundation of Korea. At Rensselaer, Dordick and Linhardt were joined in the research by Paul S. Kwon, Seok-Joon Kwon, Weihua Jin, Fuming Zhang, and Keith Fraser, and by researchers at the Korea Research Institute of Bioscience and Biotechnology in Cheongju, Republic of Korea, and Zhejiang University of Technology in Hangzhou, China.
About Rensselaer Polytechnic Institute
Founded in 1824, Rensselaer Polytechnic Institute is America’s first technological research university. Rensselaer encompasses five schools, 32 research centers, more than 145 academic programs, and a dynamic community made up of more than 7,900 students and over 100,000 living alumni. Rensselaer faculty and alumni include more than 145 National Academy members, six members of the National Inventors Hall of Fame, six National Medal of Technology winners, five National Medal of Science winners, and a Nobel Prize winner in Physics. With nearly 200 years of experience advancing scientific and technological knowledge, Rensselaer remains focused on addressing global challenges with a spirit of ingenuity and collaboration. To learn more, please visit www.rpi.edu.
Photo Credit:
Rensselaer Polytechnic Institute
In a test of antiviral effectiveness against the virus that causes COVID-19, an extract from edible seaweeds substantially outperformed remdesivir.
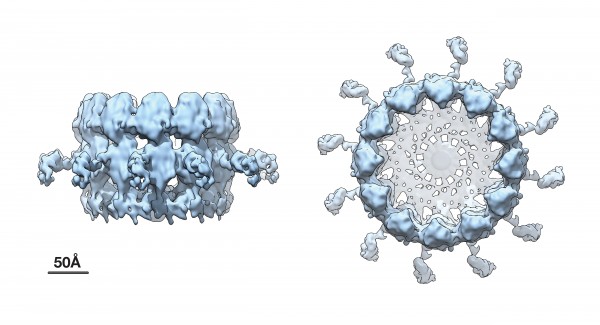
Advanced cryo-EM imaging reveals high-resolution side and top views of the viral RNA replication "crown" complex structure.
Newswise — For the first time, scientists at the Morgridge Institute for Research have generated near atomic resolution images of a major viral protein complex responsible for replicating the RNA genome of a member of the positive-strand RNA viruses, the large class of viruses that includes coronaviruses and many other pathogens.
The results should aid development of new types of antivirals and provide mechanistic insights into the virus life cycle.
“The rapidly advancing ability to visualize such crucial structures is game changing,” says Paul Ahlquist, director of the John W. and Jeanne M. Rowe Center for Virology Research at the Morgridge Institute and professor of oncology and molecular virology at the University of Wisconsin-Madison. Other authors of the study included Nuruddin Unchwaniwala, Hong Zhan, Janice Pennington, Mark Horswill and Johan den Boon.
Using an advanced technique called cryoelectron microscope (cryo-EM) tomography, Ahlquist and his team built upon their previous work, which first revealed the existence of this crown-like viral RNA replication complex.
The new research, published July 20 in the Proceedings of the National Academy of Sciences (PNAS), shows the replication crown complex at a dramatically improved resolution of approximately 8.5 angstroms, which corresponds to the spacing of a few atoms.
“Cryo-EM has recently gone through a quantum leap in its capabilities,” Ahlquist says. “In this study our research group combined multiple advances to greatly improve sample preparation, image acquisition and image processing, and to map the position of specific protein domains in the complex.”
The positive-strand RNA viruses addressed in this work are the largest of six genetic classes of viruses and include many important pathogens such as the Zika, dengue and chikungunya viruses, as well as coronaviruses like SARS-CoV-2, cause of the current COVID-19 pandemic.
In each positive-strand RNA virus, most of the viral genes are devoted to a single process: replicating the viral RNA genome.
“Given this massive investment of resources, viral RNA genome replication is arguably one of the most important processes in infection, and It is already a major target for virus control,” Ahlquist says.
Within an infected cell, viral RNA replication occurs at modified cellular membranes, often in association with spherules, virus-induced vesicles approximately 50–100 nanometers in size. Ahlquist and his team previously showed that in each such genome replication complex, a copy of the viral RNA genome or chromosome is protected inside the spherule vesicle to function as a replication template. The replication complex repeatedly copies this archival viral RNA chromosome to produce new progeny genomes that are released through a membranous neck on the vesicle into the cytoplasm, where they are incorporated as the payload of new infectious virions.
This prior work further showed that the key viral protein that induces the replication vesicles and copies the viral RNA resides in a striking ring or crown structure that sits atop the cytoplasmic side of the spherule neck that connects with the cytoplasm.
The new higher resolution cryo-EM images and complementary results show that the crown is composed of twelve copies of the key viral RNA replication protein arranged like staves in a barrel. Additionally, the images revealed zipper-like interactions that act like hoops on a barrel to join adjacent segments together to form the ring-like crown. These zippering interactions correspond well with multimerizing interactions that the Ahlquist group has previously mapped in this protein.
The viral RNA replication protein that forms the crown is an extremely large, multi-domain, multi-functional protein, nearly 1000 amino acids in size. This protein contains RNA polymerase and RNA capping domains— two enzymatic domains that are conserved across numerous positive-strand RNA viruses for synthesizing new viral genome copies—plus other domains for multimerizing, binding membranes and other functions.
How these domains are physically organized in the crown structure is one of the most important issues for understanding how the replication complex functions, and was one of several strong motivations for defining the high-resolution crown structure.
Using an approach that combined a genetically engineered, site-specific tag with labeling by nanoscale gold particles visible in cryo-EM, the researchers found that the C-terminal polymerase end of the viral RNA replication protein is positioned at the apex of the crown, leaving the N-terminal capping domain at the bottom of the structure to interact with the membrane.
This apical position of the polymerase has important mechanistic implications for early steps in the replication process that recruit the starting viral RNA template into the complex and form the replication vesicle, as well as for later steps in which the template is copied to make new progeny genomes to be packaged into infectious virus particles. These results provide a strong foundation for further experiments to define the replication complex structure and function at even higher levels.
“We hope to continue to improve the RNA replication complex crown structure to provide additional important refinements in future,” Ahlquist says. “We also hope to address growing indications from our work that conformational changes in these proteins are critical to their multiple functions.”
“Such advances will reveal in increasing detail how these complexes assemble and operate, and thus how they might be best attacked,” he adds. “These insights should provide the basis for novel, stronger antiviral mechanisms.”
Newswise — From mRNA vaccines entering clinical trials, to peptide-based vaccines and using molecular farming to scale vaccine production, the COVID-19 pandemic is pushing new and emerging nanotechnologies into the frontlines and the headlines.
Nanoengineers at UC San Diego detail the current approaches to COVID-19 vaccine development, and highlight how nanotechnology has enabled these advances, in a review article in Nature Nanotechnology published July 15.
“Nanotechnology plays a major role in vaccine design,” the researchers, led by UC San Diego Nanoengineering Professor Nicole Steinmetz, wrote. Steinmetz is also the founding director of UC San Diego’s Center for Nano ImmunoEngineering. “Nanomaterials are ideal for delivery of antigens, serving as adjuvant platforms, and mimicking viral structures. The first candidates launched into clinical trials are based on novel nanotechnologies and are poised to make an impact.”
Steinmetz is leading a National Science Foundation-funded effort to develop—using a plant virus— a stable, easy to manufacture COVID-19 vaccine patch that can be shipped around the world and painlessly self-administered by patients. Both the vaccine itself and the microneedle patch delivery platform rely on nanotechnology. This vaccine falls into the peptide-based approach described below.
“From a vaccine technology development point of view, this is an exciting time and novel technologies and approaches are poised to make a clinical impact for the first time. For example, to date, no mRNA vaccine has been clinically approved, yet Moderna’s mRNA vaccine technology for COVID-19 is making headways and was the first vaccine to enter clinical testing in the US.”
As of June 1, there are 157 COVID-19 vaccine candidates in development, with 12 in clinical trials.
“There are many nanotechnology platform technologies put toward applications against SARS-CoV-2; while highly promising, many of these however may be several years away from deployment and therefore may not make an impact on the SARS-CoV-2 pandemic,” Steinmetz wrote. “Nevertheless, as devastating as COVID-19 is, it may serve as an impetus for the scientific community, funding bodies, and stakeholders to put more focused efforts toward development of platform technologies to prepare nations for readiness for future pandemics,” Steinmetz wrote.
To mitigate some of the downsides of contemporary vaccines—namely live-attenuated or inactivated strains of the virus itself-- advances in nanotechnology have enabled several types of next-generation vaccines, including:
Peptide-based vaccines: Using a combination of informatics and immunological investigation of antibodies and patient sera, various B- and T-cell epitopes of the SARS-CoV-2 S protein have been identified. As time passes and serum from convalescent COVID-19 patients are screened for neutralizing antibodies, experimentally-derived peptide epitopes will confirm useful epitope regions and lead to more optimal antigens in second-generation SARS-CoV-2 peptide-vaccines. The National Institutes of Health recently funded La Jolla Institute for Immunology in this endeavor.
Peptide-based approaches represent the simplest form of vaccines that are easily designed, readily validated and rapidly manufactured. Peptide-based vaccines can be formulated as peptides plus adjuvant mixtures or peptides can be delivered by an appropriate nanocarrier or be encoded by nucleic acid vaccine formulations. Several peptide-based vaccines as well as peptide-nanoparticle conjugates are in clinical testing and development targeting chronic diseases and cancer, and OncoGen and University of Cambridge/DIOSynVax are using immunoinformatics-derived peptide sequences of S protein in their COVID-19 vaccine formulations.
An intriguing class of nanotechnology for peptide vaccines is virus like particles (VLPs) from bacteriophages and plant viruses. While non-infectious toward mammals, these VLPs mimic the molecular patterns associated with pathogens, making them highly visible to the immune system. This allows the VLPs to serve not only as the delivery platform but also as adjuvant. VLPs enhance the uptake of viral antigens by antigen-presenting cells, and they provide the additional immune-stimulus leading to activation and amplification of the ensuing immune response. Steinmetz and Professor Jon Pokorski received an NSF Rapid Research Response grant to develop a peptide-based COVID-19 vaccine from a plant virus: https://jacobsschool.ucsd.edu/news/news_releases/release.sfe?id=3005. Their approach uses the Cowpea mosaic virus that infects legumes, engineering it to look like SARS-CoV-2, and weaving antigen peptides onto its surface, which will stimulate an immune response.
Their approach, as well as other plant-based expression systems, can be easily scaled up using molecular farming. In molecular farming, each plant is a bioreactor. The more plants are grown, the more vaccine is made. The speed and scalability of the platform was recently demonstrated by Medicago manufacturing 10 million doses of influenza vaccine within one month. In the 2014 Ebola epidemic, patients were treated with ZMapp, an antibody cocktail manufactured through molecular farming. Molecular farming has low manufacturing costs, and is safer since human pathogens cannot replicate in plant cells.
Nucleic-acid based vaccines: For fast emerging viral infections and pandemics such as COVID-19, rapid development and large scale deployment of vaccines is a critical need that may not be fulfilled by subunit vaccines. Delivering the genetic code for in situ production of viral proteins is a promising alternative to conventional vaccine approaches. Both DNA vaccines and mRNA vaccines fall under this category and are being pursued in the context of the COVID-19 pandemic.
DNA vaccines are made up of small, circular pieces of bacterial plasmids which are engineered to target nuclear machinery and produce S protein of SARS-CoV-2 downstream.
mRNA vaccines on the other hand, are based on designer-mRNA delivered into the cytoplasm where the host cell machinery then translates the gene into a protein – in this case the full-length S protein of SARS-CoV-2. mRNA vaccines can be produced through in vitro transcription, which precludes the need for cells and their associated regulatory hurdles
While DNA vaccines offer higher stability over mRNA vaccines, the mRNA is non-integrating and therefore poses no risk of insertional mutagenesis. Additionally, the half-life, stability and immunogenicity of mRNA can be tuned through established modifications.
Several COVID-19 vaccines using DNA or RNA are undergoing development: Inovio Pharmaceuticals has a Phase I clinical trial underway, and Entos Pharmeuticals is on track for a Phase I clinical trial using DNA. Moderna’s mRNA-based technology was the fastest to Phase I clinical trial in the US, which began on March 16th, and BioNTech-Pfizer recently announced regulatory approval in Germany for Phase 1/2 clinical trials to test four lead mRNA candidates.
Subunit vaccines: Subunit vaccines use only minimal structural elements of the pathogenic virus that prime protective immunity-- either proteins of the virus itself or assembled VLPs. Subunit vaccines can also use non-infectious VLPs derived from the pathogen itself as the antigen. These VLPs are devoid of genetic material and retain some or all of the structural proteins of the pathogen, thus mimicking the immunogenic topological features of the infectious virus, and can be produced via recombinant expression and scalable through fermentation or molecular farming. The frontrunners among developers are Novavax who initiated a Phase I/II trial on May 25, 2020. Also Sanofi Pasteur/GSK, Vaxine, Johnson & Johnson and the University of Pittsburgh have announced that they expect to begin Phase I clinical trials within the next few months. Others including Clover Biopharmaceuticals and the University of Queensland, Australia are independently developing subunit vaccines engineered to present the prefusion trimer confirmation of S protein using the molecular clamp technology and the Trimer-tag technology, respectively.
Delivery device development
Lastly, the researchers note that nanotechnology’s impact on COVID-19 vaccine development does not end with the vaccine itself, but extends through development of devices and platforms to administer the vaccine. This has historically been complicated by live attenuated and inactivated vaccines requiring constant refrigeration, as well as insufficient health care professionals where the vaccines are needed.
“Recently, modern alternatives to such distribution and access challenges have come to light, such as single-dose slow release implants and microneedle-based patches which could reduce reliance on the cold chain and ensure vaccination even in situations where qualified health care professionals are rare or in high demand,” the researchers write. “Microneedle-based patches could even be self-administered which would dramatically hasten roll-out and dissemination of such vaccines as well as reducing the burden on the healthcare system.”
Pokorski and Steinmetz are co-developing a microneedle delivery platform with their plant virus COVID-19 vaccine for both of these reasons.
This work is supported by a grant from the National Science Foundation (NSF CMMI-2027668)
“Advances in bio/nanotechnology and advanced nanomanufacturing coupled with open reporting and data sharing lay the foundation for rapid development of innovative vaccine technologies to make an impact during the COVID-19 pandemic,” the researchers wrote. “Several of these platform technologies may serve as plug-and-play technologies that can be tailored to seasonal or new strains of coronaviruses. COVID-19 harbors the potential to become a seasonal disease, underscoring the need for continued investment in coronavirus vaccines.”
Photo Credit:
David Baillot/ UC San Diego Jacobs School of Engineering
UC San Diego Nanoengineering Professor Nicole Steinmetz is using a plant virus to develop a COVID-19 vaccine.
Newswise — COLUMBUS, Ohio – A new study by researchers at The Ohio State University Wexner Medical Center and The Ohio State University College of Medicine documents a clear link between the stress hormone cortisol and higher blood sugar levels in people with type 2 diabetes.
The study published online in the journal Psychoneuroendocrinology.
“In healthy people, cortisol fluctuates naturally throughout the day, spiking in the morning and falling at night,” said Dr. Joshua J. Joseph, an endocrinologist and researcher at The Ohio State Wexner Medical Center’s Diabetes and Metabolism Research Center who led the study. “But in participants with type 2 diabetes, cortisol profiles that were flatter throughout the day, had higher glucose levels.”
Previous research has shown that stress and depression are two of the major causes of a flatter cortisol profile. These sustained levels of cortisol make it much more difficult to control blood sugar and manage the disease, which is why it is so important for those with type 2 diabetes to find ways to reduce stress.
“We have begun a new trial to examine if mindfulness practices can lower blood sugar in those with type 2 diabetes,” said Joseph, an assistant professor in the College of Medicine. “But this isn’t the only effective form of stress relief. It’s important to find something you enjoy and make it a part of your everyday routine.”
The relationship of cortisol with glucose levels was only observed in those with diabetes. However, Dr. Joseph and his team believe the stress hormone likely plays an important role in diabetes prevention and they continue to research the connection between cortisol and the development of diabetes and cardiovascular disease.
More than 30 million Americans have Type 2 diabetes, according to the Centers for Disease Control and Prevention. With Type 2 diabetes, your body doesn’t use insulin properly. Some people can manage their blood sugar levels with healthy eating and exercise, while others may need medication or insulin to help manage it.
“Most people with Type 2 diabetes know the importance of exercising regularly, eating a healthy diet, and getting plenty of rest. But stress relief is a crucial and often forgotten component of diabetes management,” Joseph said. “Whether it’s a yoga class, taking a walk or reading a book, finding ways to lower your stress levels is important to everyone’s overall health, especially for those with type 2 diabetes.”
Joseph collaborated with researchers at Johns Hopkins University; University of Maryland School of Medicine; University of California, Los Angeles; University of Michigan; Boston University and the National Institute on Aging, National Institutes of Health.
Photo credit: The Ohio State University Wexner Medical Center
A persistent cough and fever have been confirmed as the most prevalent symptoms associated with COVID-19 according to the new international research
Data analysed from over 24,000 patients from nine countries including the UK, China and the US
University of Sheffield medical student helped conduct the study, which is one of the biggest reviews (or meta data analysis) into COVID-19 symptoms
Newswise — A persistent cough and fever have been confirmed as the most prevalent symptoms associated with COVID-19, according to a major international review of patient data.
Other major symptoms include fatigue, losing the ability to smell and difficulty in breathing.
The study ratifies the list of symptoms compiled by the World Health Organisation at the start of the pandemic.
A team of researchers including a final year medical student from the University of Sheffield combined data from 148 separate studies, to identify the common symptoms experienced by more than 24,000 patients from nine countries including the UK, China and the US.
The study - published in the online journal PLoS One - is one of the biggest reviews, or meta analysis, ever conducted into COVID-19 symptoms.
Of the 24,410 cases, the study found:
*78 per cent had a fever. Although this tended to vary across countries, with 72 per cent of fever reported by patients in Singapore and 32 per cent in Korea.
*57 per cent reported a cough. Again, this varied across countries, with 76 per cent of patients reporting a cough in the Netherlands compared to 18 per cent in Korea.
*31 per cent said they had suffered fatigue.
*25 per cent lost the ability to smell.
*23 per cent reported difficulty breathing.
The researchers believe the variation in the prevalence of symptoms between countries is due, in part, to the way data was collected.
Of those patients who needed hospital treatment, 17 per cent needed non-invasive ventilation; 19 per cent had to be looked after in an intensive care unit, nine per cent required invasive ventilation and two per cent needed extracorporeal membrane oxygenation (an artificial lung).
Michael Grant, a final year medical student from the University of Sheffield, said: “The impact of COVID-19 on daily activities has been immense.
“An accurate estimation of symptom prevalence, as provided by this study, is essential to combat COVID-19”
“As a medical student, it has been a tremendous opportunity and a great privilege to contribute to research that will be used to inform public health opinion.”
Mr Ryckie Wade, a surgeon and Clinical Research Fellow at the Leeds Institute of Medical Research, who supervised the research, said: "This analysis confirms that a cough and fever were the most common symptoms in people who tested positive with COVID-19.
"This is important because it ensures that people who are symptomatic can be quarantined.
"The study gives confidence to the fact that we have been right in identifying the main symptoms and it can help determine who should get tested."
The study involved academics from the University of Leeds with colleagues from the University of Sheffield, University of Bristol, Imperial College, London, and the Belgium Cancer Centre. The research was funded by the UK's National Institute for Health Research and VALCOR, in Belgium. The researchers acknowledge there are people who had the virus but did not display any symptoms.
Additional information
The University of Sheffield’s Medical School trains world class clinical and researchers who prolong and improve lives everyday.
The vision of the Medical School is to achieve research excellence and impact by translating scientific discoveries into meaningful advances that can benefit patients and the general population. This vision is supported by competitive funding from major national and international research bodies including Government, Research Councils, industry and charities. For more information please visit: https://www.sheffield.ac.uk/medicine
The University of Sheffield
With almost 29,000 of the brightest students from over 140 countries, learning alongside over 1,200 of the best academics from across the globe, the University of Sheffield is one of the world’s leading universities.
A member of the UK’s prestigious Russell Group of leading research-led institutions, Sheffield offers world-class teaching and research excellence across a wide range of disciplines.
Unified by the power of discovery and understanding, staff and students at the university are committed to finding new ways to transform the world we live in.
Sheffield is the only university to feature in The Sunday Times 100 Best Not-For-Profit Organisations to Work For 2018 and for the last eight years has been ranked in the top five UK universities for Student Satisfaction by Times Higher Education.
Sheffield has six Nobel Prize winners among former staff and students and its alumni go on to hold positions of great responsibility and influence all over the world, making significant contributions in their chosen fields.
Global research partners and clients include Boeing, Rolls-Royce, Unilever, AstraZeneca, Glaxo SmithKline, Siemens and Airbus, as well as many UK and overseas government agencies and charitable foundations.
A new study reveals that the common asthma drug salbutamol may offer potential as a treatment for Alzheimer's disease
Newswise — A new study reveals that the common asthma drug salbutamol may offer potential as a treatment for Alzheimer's disease.
Alzheimer's disease is the most common form of dementia, affecting 47 million people worldwide and its prevalence is expected to triple to more than 130 million cases by 2050.
No effective treatments that cure the disease or slow down its progression have been discovered. However, this new early-stage study has revealed that repurposing an existing drug, salbutamol, offers significant potential as a low cost and rapid response option.
Extensive analytical in-vitro experiments conducted by the research team show that salbutamol is effective at reducing the accumulation of insoluble fibres of the tau protein - which is found in the brains of people with Alzheimer's disease. These microscopic fibres accumulate into neurofibrillary tangles and can cause neuron destabilisation, brain cell death, and are a key characteristic of the disease's progression.
Much Alzheimer's disease research has focused on the build-up of amyloid plaques, caused by misfolding of the amyloid-β protein. However, because of disappointing results from numerous therapies targeting Aβ aggregation, more attention is shifting towards tau.
This study, led by researchers at Lancaster University, used a new automated 'high throughput' screening approach to study the structure of the misfolding tau protein with a special analytical technique called 'Synchrotron Radiation Circular Dichroism' (SRCD) at Diamond Light Source, the UK national synchrotron light source in Oxfordshire. With this powerful technique they were able to look at a selection of more than 80 existing compounds and drugs simultaneously to determine their effectiveness at preventing the formation of tau fibrils.
This method confirmed the compound epinephrine, more commonly known as adrenaline, was effective at stabilising the tau proteins and preventing the formation of tau tangles. However, our bodies do not easily absorb epinephrine and it rapidly gets metabolised, so the scientists then looked at a range of readily available compounds with similar chemical structures. This search revealed four current drugs as possible candidates - etamivan, fenoterol, dobutamine and salbutamol.
Etaminvan and fenoterol were found to have little effect on the assembly of tau tangles. Dobutamine, which is used for the rapid treatment of heart attacks and heart failure, was found to have some benefit. However, because its effects are very short-lived, and because it needs to be administered intravenously, it is not ideal as a basis for treatment of Alzheimer's disease.
Further tests using a range of analytical techniques all revealed salbutamol could inhibit tau aggregation in vitro. Tests where salbutamol was added to solutions containing tau resulted in drastically reduced density of fibrous tau structures responsible for the tau neurofibrillary tangles.
The researchers believe that salbutamol interacts with an early stage of tau fibril formation, reducing their ability to form an initial nucleus which drives the aggregation process.
Because it is easily ingested, absorbed into the brain, and remains in the body for several hours, salbutamol has attractive properties as a research avenue for potential new treatment for Alzheimer's.
Dr David Townsend, of Lancaster university and lead author of the research, said: "Our work highlights the potential impact of repurposing drugs for secondary medical uses, by discovering a novel therapeutic strategy that impedes the molecular pathology of Alzheimer's disease, and which may have otherwise gone unstudied.
"Salbutamol has already undergone extensive human safety reviews, and if follow up research reveals an ability to impede Alzheimer's disease progression in cellular and animal models, this drug could offer a step forward, whilst drastically reducing the cost and time associated with typical drug development."
Professor David Middleton, co-author of the research, said: "This work is in the very early stages and we are some way from knowing whether or not salbutamol will be effective at treating Alzheimer's disease in human patients. However, our results justify further testing of salbutamol, and similar drugs, in animal models of the disease and eventually, if successful, in clinical trials."
Dr Rohanah Hussain, of Diamond Light Source, Senior Beamline Scientist and co-author said: "Diamond B23 beamline unique micro-collimated beam has made high throughput CD possible allowing the screening of many compounds through structural activity correlation crucial in drug discovery."
The researchers say that current asthma inhalers result in only a small amount of salbutamol reaching the brain and so, if further research is successful, a new delivery method would also need to be developed. They add that future research could also focus on other asthma drugs that are chemically similar to salbutamol, but which circulate in the bloodstream for much longer.
Olafur Palsson, PsyD, professor of medicine in the UNC School of Medicine, led a study investigating the emotional and mental impact of the COVID-19 pandemic in the U.S. The study authors are releasing their findings directly to the public in an effort to provide guidance and understanding to individuals and clinicians as the pandemic evolves.
Newswise — CHAPEL HILL, NC – The findings of a nationwide survey assessing the effects of the COVID-19 pandemic on the emotional wellbeing of the U.S. adult population have been released online. The survey was a collaboration between UNC School of Medicine and Harvard Medical School, and was quickly organized to gain an understanding of how individuals are responding to the stressors of isolation and quarantine, record unemployment levels, and the virus’ threat to their health.
“We were very careful to balance the demographics and geographic distribution of survey respondents so that it is an accurate representation of the U.S., and essentially a snapshot of the population,” said principal investigator Olafur Palsson, PsyD, professor of medicine in the division of gastroenterology and hepatology in the UNC School of Medicine.
The study consisted of a nationwide internet survey of 1,500 people conducted during the second half of May, when the pandemic was just beyond its peak in the nation. It is worth noting that the survey ended on May 30, five days after George Floyd was killed by a police officer in Minnesota. Nearly 90 percent of survey responses were collected before the movement across the U.S. to increase recognition of systemic racism.
“We saw that levels of concern or worry differed among different demographics including age and race,” Palsson said. “Women and men tended to experience similar degrees of emotional impact associated with the pandemic on average, but younger people were more affected by it than older individuals, and the Hispanic population seemed to be the most impacted emotionally when comparing ethnic groups.”
The study assessed a broad range of specific emotional effects related to the pandemic, and found that there were stressors that seemed to affect a majority of the population. Nearly 80 percent of respondents were frustrated on some level with not being able to do what they normally enjoy doing. Around the same number were worried about their own health, and nearly 90 percent of those surveyed were more worried about the health of loved ones than before the COVID-19 pandemic.
“Most people have family or friends with chronic illness or health conditions that would make them more vulnerable to the virus, so worry about loved ones seems to be a shared feeling no matter who you are,” Palsson said.
The emotional and mental impact of the pandemic could have long-term implications on well-being. Palsson and co-authors Sarah Ballou, PhD, instructor in medicine at Harvard Medical School and director of gastrointestinal psychology at Beth Israel Deaconess Medical Center, and Sarah Gray, PsyD, instructor in psychology at Harvard Medical School and practicing psychologist at Massachusetts General Hospital, wanted to get the results of the study out as quickly as possible to aid in addressing what could be a growing mental health crisis.
“Understanding the psychological side of this pandemic has been somewhat neglected because there is so much else that is of concern right now,” Palsson said. “Our survey findings indicate that the anxiety and depression related to the emotional impact of these events are pushing more and more people into the clinical category of what is diagnosable as a mental health condition. We want people to seek the help they need.”
A list of resources is available on the study’s website (http://pandemicimpactreport.com/index.html), along with a downloadable version of the study’s findings.
Photo credit: UNC School of Medicine
Olafur Palsson, PsyD
Newswise — TALLAHASSEE, Fla. — Social distancing during the COVID-19 pandemic has not led to an overall increase in loneliness among Americans.
That’s the takeaway from a comprehensive, nationwide study by Florida State University College of Medicine researchers who surveyed more than 2,000 people before and during the enactment of stay-at-home policies in response to the COVID-19 pandemic.
The study on how loneliness and perceived support has changed from before to during the pandemic is published in American Psychologist.
“There has been a lot of worry that loneliness would increase dramatically because of the social distancing guidelines and restrictions,” said lead author Martina Luchetti, an assistant professor at the College of Medicine. “Contrary to this fear, we found that overall loneliness did not increase. Instead, people felt more supported by others than before the pandemic. Even while physically isolated, the feeling of increased social support and of being in this together may help limit increases in loneliness.”
The paper is part of a larger study College of Medicine researchers are doing on COVID-19 to look at changes in mental health during the COVID-19 crisis and how psychological factors contribute to various aspects of response to the pandemic.
The study involved a nationwide panel of adults ages 18 to 98. Participants first completed a survey in early February unrelated to COVID-19, before the virus was widely known to be a threat to the U.S.
As the threat was being realized, researchers contacted participants again for two more surveys — one in mid-March during the 15-day period to slow the spread based on White House guidelines and another in late April as the guidelines were about to expire. More than 2,000 responses to the surveys were included in the findings.
The study also looked for increased loneliness in specific at-risk groups, finding only modest evidence of a small increase in loneliness among older adults. Older adults reported less loneliness overall compared to younger age groups, despite an increase in loneliness during the acute phase of the outbreak. That increase in feelings of loneliness among older adults was temporary, leveling off after the issuance of stay-at-home orders.
Individuals living alone and those with at least one chronic condition reported feeling lonelier at the outset but did not increase in loneliness after social-distancing measures were implemented.
“Despite a small increase among some individuals, we found overall remarkable resilience in response to COVID-19,” said Angelina Sutin, associate professor of Behavioral Sciences and Social Medicine and senior author.
Loneliness already was a known health risk before the pandemic, and it has been linked to increased risks of morbidity and mortality. Surveys have found that 35 percent of adults 45 and older report feeling lonely and 43 percent of those over 60 report experiencing loneliness at least some of the time.
Some studies suggest that loneliness is even more pervasive among younger adults.
“In the context of the coronavirus pandemic, it may be particularly difficult to reconnect with others given the restrictions on in-person social gatherings,” Luchetti said. “Even these transient feelings of loneliness can have a negative effect on health, meaning there could be dangerous unintended consequences if loneliness increases in response to the restrictive measures taken as a result of the pandemic.”
Yet from the start of the pandemic, there have been anecdotal reports of people calling their family and friends more often and finding creative ways to stay connected. This outpouring of support may have shielded them from potential increases in loneliness.
The pandemic is also something that everybody is going through.
“Just knowing that you are not alone and that everyone is going through the same restrictions and difficulties may be enough in the short term to keep feelings of loneliness down,” Sutin said.