News
MOUNT SINAI DEVELOPS A SAFE, LOW-COST COVID-19 VACCINE - COULD HELP LOW- AND MIDDLE-INCOME COUNTRIES
Newswise — A low-cost COVID-19 vaccine candidate that could be produced in the United States and worldwide using existing influenza virus manufacturing infrastructure has been developed by researchers at Icahn School of Medicine at Mount Sinai, with the potential to rapidly produce hundreds of millions of vaccine doses to mitigate the impact of the current pandemic and future viral outbreaks. The vaccine, which works the same way many flu vaccines do, is undergoing clinical development planning in several countries (including Mexico) through a licensing agreement with Mount Sinai. Early-stage clinical trials are also underway at Mount Sinai Health System in New York.
“To contain the spread of the virus worldwide, a vaccine that is both effective and cost-effective is urgently needed, especially in low- and middle-income countries with limited resources,” says Peter Palese, MD, Horace W. Goldsmith Professor and Chair of Microbiology at Icahn Mount Sinai, and senior author of two studies examining the effects of so-called Newcastle disease virus (NDV) vaccines in animal models (EBioMedicine, November 2020, and Vaccines, December 2020). “Our work suggests that an NDV-based vaccine, which can be produced from embryonated chicken eggs, would be a safe and highly scalable way to meet the vast demands of the global vaccine market.”
Most COVID-19 vaccines work by exposing people to the “spike” protein, an important part of the structure of SARS-CoV-2, the virus that causes COVID-19. But they differ in how they get the spike protein into the recipient. Mount Sinai’s vaccine works by introducing the spike protein into the body via the harmless NDV virus, prompting the body’s cells to make copies of the spike protein. When this occurs, the immune system begins to produce antibodies and T cells to specifically target the spike protein; these will neutralize any foreign intruder, like SARS-CoV-2, that contains it.
The NDV vaccine against SARS-CoV-2 was created by a globally recognized team of virologists from Mount Sinai, including Dr. Palese; Florian Krammer, PhD, Mount Sinai Professor in Vaccinology; and Adolfo Garcia-Sastre, PhD, Irene and Dr. Arthur M. Fishberg Professor of Medicine and Director of the Global Health and Emerging Pathogens Institute at Mount Sinai. “Our study demonstrated that the neutralizing antibodies produced by the NDV vaccine provided strong protection in animal models from SARS-CoV-2 infection,” notes Dr. Krammer. “Also, because the NDV is not a human pathogen, the spike antigen could be delivered more efficiently and without being compromised by any pre-existing immunity in humans. Another advantage is the fact that NDV-based vaccines have been tested extensively in human trials and have compiled a very good safety record over the years.”
In Mexico, a licensing agreement between Mount Sinai and Laboratorio Avi-Mex S.A. de C.V. (Avimex), a veterinary pharmaceutical company, will enable that country to soon begin Phase 1 trials in humans of a COVID-19 vaccine using NDV vector technology. Avimex began collaborating with Mount Sinai in 2003 on developing veterinary influenza vaccines and has since produced millions of doses based on the NDV vector platform.
“Being able to vaccinate populations in all parts of the world, and not just those in high-income countries, is critical if we’re going to establish herd immunity and contain the spread of COVID-19,” says Dr. Palese. “We believe that NDV-based vaccines can be a vital part of the solution. They could lead to vaccination of a large percentage of the world’s population over a very short period by using existing technology and infrastructure in a highly cost-effective, efficient, and safe way.”
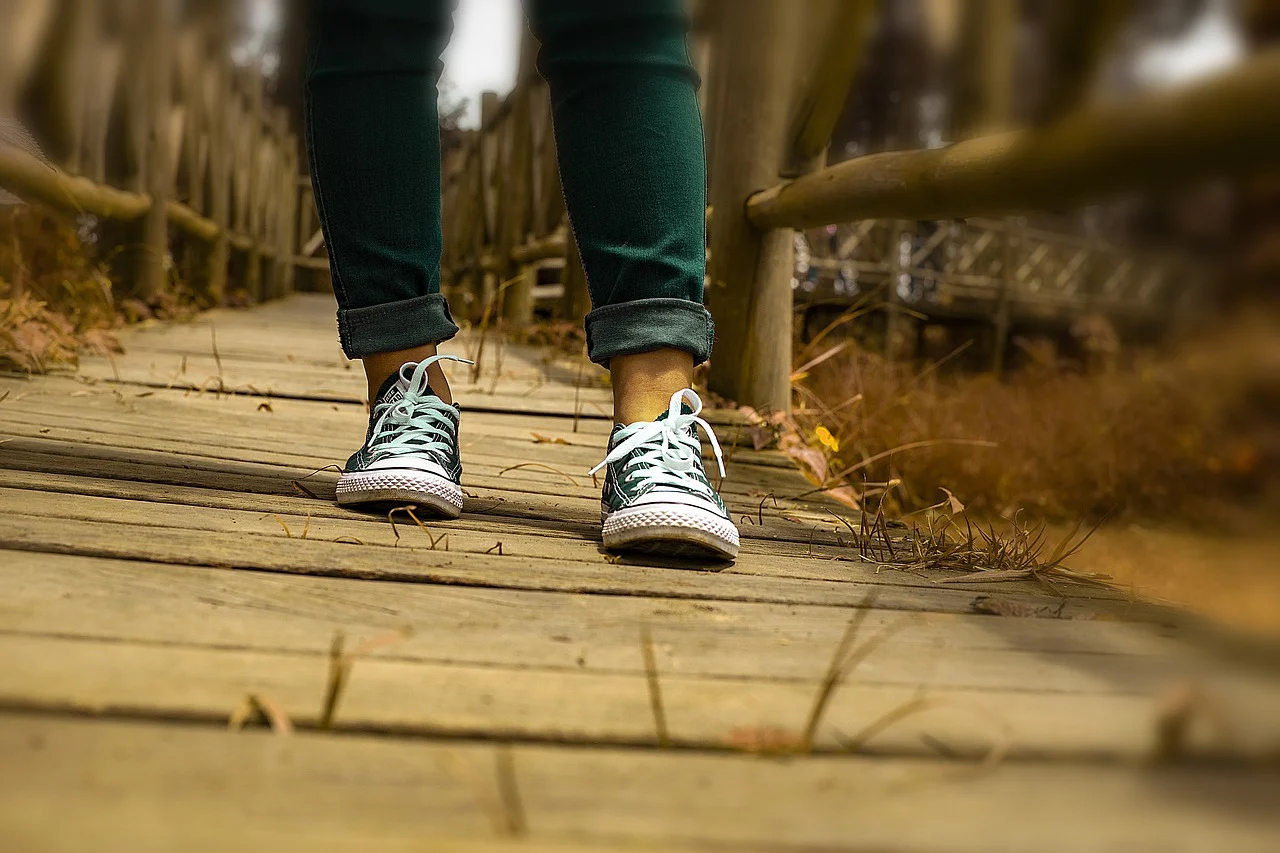
Newswise — A new study led by Washington University School of Medicine in St. Louis and the National Cancer Institute (NCI) has identified an association between slow walking pace and an increased risk of death among cancer survivors.
While the study does not establish that slow walking is a cause of death, the association persisted across at least nine tumor types. Investigators now call for more research into these relationships and whether targeted interventions such as physical activity programs could help cancer survivors improve their ability to walk and increase survival after cancer diagnosis and treatment.
The study, a collaboration between Washington University, the NCI of the National Institutes of Health (NIH), the University of North Carolina and George Washington University, appears March 4 in Cancer Epidemiology, Biomarkers & Prevention, a journal of the American Association for Cancer Research.
“Cancer survivors are living longer than ever – and that’s good news,” said first author Elizabeth A. Salerno, PhD, an assistant professor of surgery in the Division of Public Health Sciences at Washington University. “But it’s important to improve our understanding of how the diagnosis and treatment of a broad range of cancers may affect walking pace during survivorship — a potentially modifiable risk factor — which could lead to new treatment and rehabilitation strategies to improve the health of these patients.”
The researchers studied over 233,000 participants enrolled in the National Institutes of Health-American Association of Retired Persons (NIH-AARP) Diet and Health Study. Participants, who were ages 50 to 71, answered questionnaires about their overall health and walking pace, and whether they had any disability related to walking, such as walking at a very slow pace or being unable to walk. After the assessment, participants were followed for several years.
Compared with healthy controls enrolled in the study, cancer survivors were 42% more likely to report walking at the slowest pace and 24% more likely to report being disabled. Among cancer survivors, those who walked at the slowest pace had more than twofold increased risk of death from any cause, compared with those reporting the fastest walking pace.
The association between the slowest walking pace and a significantly increased risk of death from any cause held for nine cancer types, including breast, colon, melanoma, Non-Hodgkin lymphoma, oral, prostate, rectal, respiratory and urinary cancers. The association between mobility disability (not just slow pace) and death was even stronger and included all nine of the cancers mentioned above, plus endometrial, endocrine, ovarian and stomach cancers.
While slow walking pace also was linked to increased mortality that was due to any cause among individuals without a cancer diagnosis, the risk of death more than doubled for cancer survivors. Compared with individuals without a cancer diagnosis who walked at the fastest pace, cancer survivors who walked the slowest had more than tenfold increased risk of death from any cause. Cancer survivors with mobility disability had more than fivefold increased risk of death compared with individuals with no cancer diagnosis or disability.
The researchers noted that cancer survivors reported difficulties walking five years or more after cancer diagnosis and treatment, suggesting that the detrimental effects of cancer diagnosis and therapy are widespread across cancer types and long lasting, creating opportunities for intervening to help such patients improve their walking ability and pace.
“To our knowledge, this analysis is the first to explore the relationship between cancer, walking pace and subsequent mortality in 15 different cancer types,” said Salerno, who conducted this research while a postdoctoral researcher at the NCI. “Next steps include identifying the underlying reasons for these associations. It’s possible that slow walking may be due to the cancer itself, adverse effects of treatment, or changes in lifestyle. There is still much to be learned about these complex relationships, but our results highlight the importance of monitoring and even targeting walking pace after cancer.”
Study published in American Journal of Critical Care reports how five years of rapid mortality reviews of 500+ medical ICU patient deaths sparked systemic changes, quality improvements
Newswise — An in-person multidisciplinary rapid mortality review (RMR) process helped identify specific areas to improve patient care at a Los Angeles hospital, according to a study published in American Journal of Critical Care (AJCC).
The novel approach helped front-line clinicians understand both individual- and systems-level issues that contribute to mortality, with the ultimate aim of optimizing the delivery of patient care.
“Rapid Mortality Review in the Intensive Care Unit: an In-Person Multidisciplinary Improvement Initiative” explores the data generated from five years of reviewing patient deaths that occurred in the 24-bed medical intensive care unit (ICU) at Ronald Reagan University of California Los Angeles (UCLA) Medical Center.
The analysis found that the RMR process not only identified immediate concerns related to patient care but also yielded valuable insights on potentially preventable patient deaths and areas for hospital improvement initiatives.
First author Kristin Schwab, MD, is a pulmonologist and critical care physician at UCLA Health and a clinical instructor in the Division of Pulmonary and Critical Care Medicine, Department of Medicine, at UCLA’s David Geffen School of Medicine.
“Our findings suggest that these short and timely in-person meetings can be a powerful tool for efforts to both improve quality and prevent mortality in the ICU,” she said. “Bringing members of the multidisciplinary care team together for regular face-to-face discussions provided a forum that revealed concerns and solicited tangible ideas for solutions.”
Retrospective case reviews, provider surveys, and structured morbidity and mortality conferences can be useful tools for discussions about safety and quality issues, but these common tactics are unlikely to provide an efficient and practical means of reviewing all patient deaths.
The RMR process began as a pilot in 2013, with a subset of patients who had died in the medical ICU during the prior week. The subset gradually grew and by 2017, the team was attempting, during the weekly meetings, to review every death that occurred in the unit. During the five-year period, the RMR team reviewed 542 deaths, which was more than 80% of all deaths that occurred in the unit.
For each patient death, a facilitator reviewed the patient’s chart prior to the meeting to prepare to lead a semistructured interview with the care team. Questions included, “Was the death potentially preventable?” and “Are there any aspects of care that could have been improved?” followed by additional open-ended questions.
Following the meeting, the facilitator recorded a summary of the discussion into a database. The quality team reviewed the data from each meeting and referred any action items to the appropriate department.
Only 7% of deaths were deemed potentially preventable, as determined by the treatment team, RMR facilitator or both. However, the treatment team believed that care could have been improved in more than 40% of the deaths, while the facilitator identified areas for improvement in more than half of the cases.
Cases in which the patient required resuscitation after an in-hospital cardiac arrest or those in which the patient was not receiving comfort care at the time of death were associated with a higher likelihood of generating an action item.
Issues included concerns with communication or teamwork, advance care planning, delays in care, medical errors, procedural complications and hospital-acquired infections. Systems-related action items addressed lack of protocols, resource availability and throughput.
Among the identified action items, more than one in 10 led to tangible systemic change, with 29 discrete changes occurring during the study period.
Examples of completed action items include creating a standardized checklist for inbound patient transfers and changing the electronic health record to separate one-time orders from continuing orders.
To access the article and full-text PDF, visit the AJCC website at www.ajcconline.org.
Newswise — Curtin University research has found people grieving a COVID-related death would benefit from timely support and care to reduce the high risk of experiencing problems in important areas of everyday life.
Published in Journal of Pain and Symptom Management, the study is the first to focus on psychological factors that explain why people bereaved by COVID-19 might experience challenges in important areas of life, work, leisure, and relationships.
Lead author, Associate Professor Lauren Breen from the Curtin School of Population Health worked with American researchers to survey people in the United States who had lost a close person due to COVID-19 and found key psychological factors such as separation distress, dysfunctional grief, and post-traumatic stress explained why they were having trouble coping in key areas of life.
"Existing research shows that grief from deaths during the pandemic was felt more acutely than that following both deaths before the pandemic, and deaths from other natural causes," Associate Professor Breen said.
"This exacerbation of grief is due to the necessary restrictions that affect people's access to dying loved ones, limit their participation in important rituals like funerals, and reduce the physical social support they would otherwise receive from friends and family."
"There is a real need for strategies such as the integration of psychological care into palliative care to facilitate efficient and cost-effective means of supporting people who are grieving," Associate Professor Breen said.
"Better screening and assessment of bereaved people is also required, along with more accessible support services, development of improved therapies and grief interventions, and an increased number of grief specialists in the workforce."
Newswise — New Brunswick, N.J February 22, 2021 – The most frequently mutated gene in human cancers is called p53. Patients with Li-Fraumeni syndrome, which is a rare disorder that increases the risk of developing several types of cancer, often have an increased risk to develop cancers at early ages if they inherit p53 mutations. Recent studies suggest that some individuals with inherited p53 mutations do not have the early onset or high frequency of cancers, suggesting that other genetic, environmental, immunological, epigenetic, or random factors play a part in the development of cancers.
A recent study from Rutgers Cancer Institute of New Jersey tested this possibility by analyzing tumor formation and p53 mutations in mice from different genetic backgrounds. Observations from this work may further elucidate the diversity of cancers in different Li-Fraumeni patients. Senior and corresponding author of the work Wenwei Hu, PhD, researcher at Rutgers Cancer Institute and professor of radiation oncology at Rutgers Robert Wood Johnson Medical School, along with lead and corresponding author Chang S. Chan, PhD, researcher at Rutgers Cancer Institute and associate professor of medicine at Rutgers Robert Wood Johnson Medical School, share more about the findings published in Life Science Alliance (http://doi.org/10.26508/lsa.202000952).
Why is this topic important to explore?
Mutations in the p53 gene are the single most common spontaneous genetic alterations observed in human cancers. Approximately one in 20,000 individuals inherit heterozygous p53 mutations, resulting in early onset and high frequency of cancers in each patient over a lifetime. Individuals with an inherited p53 mutation have a much higher risk compared to the general population of developing adrenal cortical carcinoma, choroid plexus carcinoma, medullary blastoma, rhabdomyosarcoma and osteogenic sarcoma. There is also a high relative risk of developing breast cancer, lipomas and liposarcomas, and leiomyosarcomas. However, even within family members who share the same p53 mutation, there is great variability in what cancer types they get and when they get it, thus, it is important to explore the influence of genetics and non-genetic factors on tumor formation and tumor type. These may include environment, immunological or random factors.
Describe the work and tell us what the team discovered.
We created seven sets of mice with different genetic backgrounds, all having the same p53 mutation. These mice are prone to developing a variety of tumor types because of the p53 mutation they harbor. The tumor types these mice develop are very similar to human Li-Fraumeni patients. The mice from each genetic background are almost genetically identical and the environments are controlled to be the same. This allows us to compare the variability of the tumors within genetically identical mice to mice with different genetic backgrounds, and thus tease apart the contribution of genetics and randomness to tumor formation. We discovered that certain genetic backgrounds greatly increase the chance of developing specific tumor types and the number of tumors in a single mouse. The age at which a tumor occurs is correlated with the tissue type of that tumor, although identical tumor tissue types can occur at very different ages. Sex of the mice also impact the risk for cancer in certain genetic backgrounds. These observations present evidence for both genetic and random effects upon tumor formation in diverse groups of mice. This helps to explain the great diversity of cancers in different Li-Fraumeni patients over their lifetimes.
What are the implications of these findings?
Although the results are consistent with a series of genetic modifiers that influence the age of onset of a tumor and the tumor tissue type, the results also support random factors playing a role in the development of tumors. The most obvious random event is a spontaneous mutation in one of the many different tissue specific stem cells of the body that increase cancer risk. Other random factors may include different microbiomes from mouse to mouse, random errors in development and the adaptive immune system which is different between identical strains of mice or identical twins.
The approach in this work can lead to the identification of the gene or genes that predispose individuals to early onset tumors, the selection of the tissue type of a tumor, and enhancement of tumor risk. Genome sequencing of these tumors will help identify the genes whose mutations act with p53 mutations to influence benign and malignant tumors.
Along with Drs. Hu and Chan, other authors include Yvonne Sun, PMV Pharma; Hua Ke, Yuhan Zhao, Merzu Belete, Cen Zhang and Zhaohui Feng, Rutgers Cancer Institute; and Arnold J. Levine, Simons Center for Systems Biology, Institute for Advanced Study in Princeton.
This research was supported by grants from the National Institutes of Health, National Cancer Institute (P01CA087497-18, R01CA203965) and Department of Defense (W81XWH-18-10238). Other acknowledgements, author disclosures and other information can be found here.
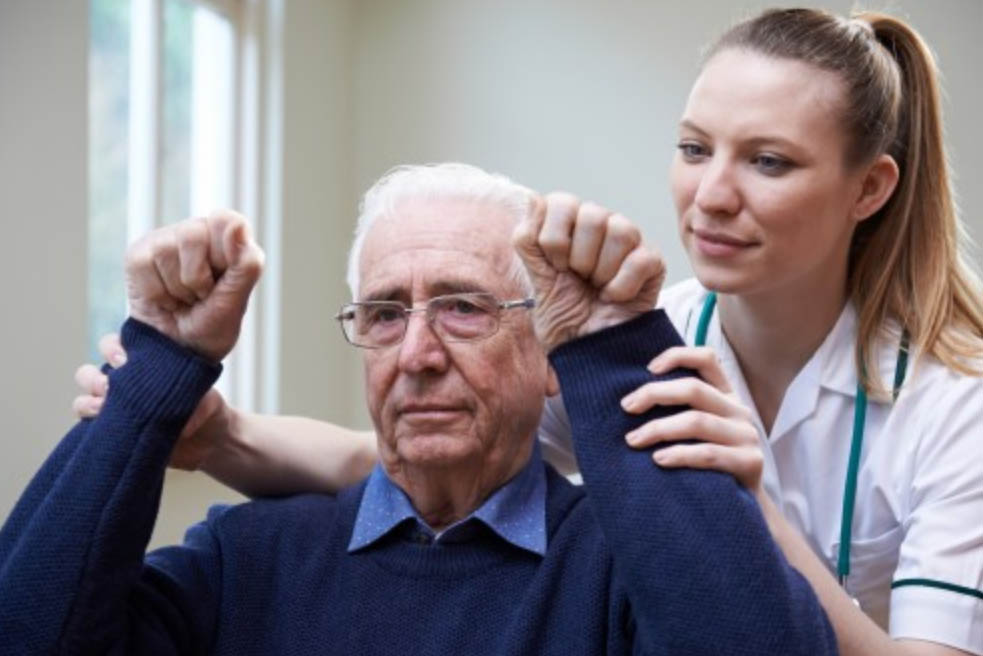
Newswise — An international study has shown, for the first time, that the capacity of the human brain to recover and rewire itself peaks around two weeks after a stroke and diminishes over time.
The finding, published today in the Neurorehabilitation and Neural Repair journal, is the result of a study in London and Adelaide that followed the recovery of 60 stroke patients up to one year after their stroke.
Lead author Dr Brenton Hordacre, from the University of South Australia, says the multi-site study showed conclusive evidence that the brain only has a small window of opportunity to more easily repair itself after stroke.
“Earlier animal studies suggested this was the case, but this is the first time we have conclusively demonstrated this phenomenon exists in humans,” Dr Hordacre says.
The researchers scanned the brains of stroke survivors as they recovered over 12 months. They found that in the initial days following an ischemic stroke (caused by a blocked artery to the brain), the brain has a greater capacity to modify its neural connections and its plasticity is increased.
“It is during this early period after stroke that any physiotherapy is going to be most effective because the brain is more responsive to treatment.
“Earlier experiments with rats showed that within five days of an ischemic stroke they were able to repair damaged limbs and neural connections more easily than if therapy was delayed until 30 days post stroke.”
The researchers used continuous transcranial magnetic stimulation (cTBS) to repetitively activate different hemispheres of the motor cortex to measure brain plasticity.
The Adelaide laboratory tested the stroke damaged motor cortex, which is the main area that controls movement. The London laboratory tested the non-stroke damaged hemisphere which is also important to help recovery.
“Our assessments showed that plasticity was strongest around two weeks after stroke in the non-damaged motor cortex. Contrary to what we expected, there was no change in the damaged hemisphere in response to cTBS.”
Dr Hordacre says the findings confirm the importance of initiating therapy as soon as possible after a stroke.
Current evidence indicates that less than eight minutes of daily therapy is dedicated to upper limb recovery within the first four weeks of a stroke.
“Delivering more treatment within this brief window is needed to help people recover after stroke.
“The next step is to identify techniques which prolong or even re-open a period of increased brain plasticity, so we can maximise recovery,” Dr Hordacre says.
The paper, “Evidence for a Window of Enhanced Plasticity in the Human Motor Cortex following Ischemic Stroke” is available at: https://journals.sagepub.com/doi/full/10.1177/1545968321992330
Researchers from the following institutions were involved in the study: University of South Australia; University College London (UCL); University of Adelaide; Hospital Universitario Ramón y Cajal and Hospital Ruber Internacional in Madrid; Queen Mary University, London; the Royal London Hospital; National Hospital for Neurology and Neurosurgery, London; Murdoch University, WA; Royal Adelaide Hospital; and the Physio Clinic, Adelaide
Photo Credit:
University of South Australia
Less than eight minutes of daily therapy is dedicated to upper limb recovery within the first four weeks of a stroke.
Newswise — DALLAS – Feb. 11, 2021 – A study led by UT Southwestern has identified a mechanism that controls the activity of proteins known as chaperones, which guide proteins to fold into the right shapes. The findings, published online today in Nature Communications, could shed light on hundreds of degenerative and neurodegenerative diseases caused by protein misfolding, such as Alzheimer’s, Parkinson’s, and Huntington’s, potentially leading to new treatments for these devastating conditions.
Every protein in the body is originally produced in a linear chain, with amino acid building blocks strung together one after another. But to fulfill their roles in cells, explains study leader Lukasz Joachimiak, Ph.D., assistant professor in the Center for Alzheimer’s and Neurodegenerative Diseases at UT Southwestern, these chains need to fold into precise shapes. Chaperones help proteins accomplish this by protecting their vulnerable portions while they shift into position and steering them to adopt the proper shape.
Every cell has a variety of chaperones that recognize and act on individual protein types. However, every chaperone isn’t active all the time, Joachimiak says. Unknown regulatory mechanisms appear to control when certain chaperones step in to guide their respective proteins to fold and when they stand aside.
Joachimiak, also a member of the Peter O’Donnell Jr. Brain Institute, and his colleagues studied a family of chaperone proteins known as Hsp40s that work in combination with other chaperones known as Hsp70s. Members of these co-chaperones are involved in the proper folding of many proteins, including tau, which play a key role in causing Alzheimer’s disease when it’s misfolded.
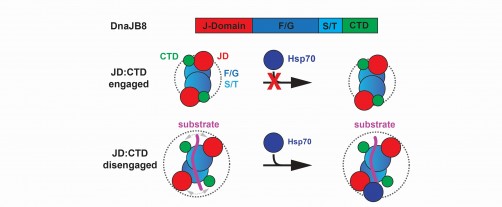
Hsp40 chaperones bind to Hsp70s through a specific portion on the Hsp40s called the J domain. But how the Hsp40s turn off this binding when it is not needed has been unclear.
To help answer this question, Joachimiak and his colleagues used a specific Hsp40 called DnaJB8 as a model. When the researchers genetically modified these proteins to glow green inside cells, they found that they didn’t just exist as individual, free-floating units – the DnaJB8 chaperones tended to form aggregates, suggesting they had some way to stick to each other. They retained this ability to agglomerate when they were isolated in petri dishes.
Using computer modeling and guided by biochemical experiments, the researchers discovered that two separate parts of this chaperone were drawn to each other through a type of chemistry called electrostatic interactions: Part of the J domain was drawn to a different part of this protein called the C-terminal domain through charged interactions. Modeling also showed that the J domain and the C-terminal domain stuck together on single molecules as well.
Joachimiak and his team validated these findings on real DnaJB8 proteins using a technique called solid-state nuclear magnetic resonance. They also showed that the J and C-terminal domains stuck to each other when they were isolated from the full DnaJB8 molecule.
The researchers suspected that the interaction between these two domains could prevent DnaJB8 from binding to its co-chaperone, an Hsp70, preventing them from jointly doing their job of guiding protein folding. Sure enough, experiments showed that the C-terminal domain of DnaJB8 competed with an Hsp70 called HspA1A when it was added to DnaJB8 in a test tube, blocking HspA1A from binding to the J domain when the C-terminal domain was bound instead.
Joachimiak notes that something may go awry in this or other regulatory mechanisms that control the activity of chaperones in protein misfolding diseases. Finding ways to control this activity through pharmaceuticals or other means could provide a new way to treat these conditions to attack the problem at its source.
“We may be able to leverage this mechanism to directly target these chaperones, activating them at will,” says Joachimiak, who is also an assistant professor of biochemistry and an Effie Marie Cain Scholar in Medical Research. “Our results could have an impact on hundreds of diseases where proteins become bad players by misfolding.”
Other UT Southwestern researchers who contributed to this study include Bryan D. Ryder, Sofia Bali, and Jaime Vaquer-Alicea.
This work was supported by a grant from The Welch Foundation (I-1928-20170325).
About UT Southwestern Medical Center
UT Southwestern, one of the premier academic medical centers in the nation, integrates pioneering biomedical research with exceptional clinical care and education. The institution’s faculty has received six Nobel Prizes, and includes 23 members of the National Academy of Sciences, 17 members of the National Academy of Medicine, and 13 Howard Hughes Medical Institute Investigators. The full-time faculty of more than 2,500 is responsible for groundbreaking medical advances and is committed to translating science-driven research quickly to new clinical treatments. UT Southwestern physicians provide care in about 80 specialties to more than 105,000 hospitalized patients, nearly 370,000 emergency room cases, and oversee approximately 3 million outpatient visits a year.
Newswise — COLUMBUS, Ohio – A new national survey of more than 2,000 Americans by The Ohio State University Wexner Medical Center finds most plan to continue many of the pandemic precautions in the name of public health, even when the pandemic is over.
As more people get vaccinated against COVID-19, there’s growing optimism for a happier and healthier future. But experts warn that life will not return to normal right away, and people should expect many health precautions and restrictions implemented during the pandemic to stick around for the foreseeable future.
“While the progress we’re making toward recovery is exciting, it’s critical that we don’t ease up on the precautions that we know have worked thus far,” said Dr. Iahn Gonsenhauser, chief quality and patient safety officer at The Ohio State Wexner Medical Center. “Masks and physical distancing are still our best weapons for limiting spread and, now that we have a vaccine, will make those precautions even more effective and will drive new cases way down if we stay the course.”
The survey found that nearly three-quarters (72%) of Americans plan to continue to wear masks in public, four out of five (80%) will still avoid crowds and 90% plan to keep up frequent handwashing and sanitizer use after COVID-19. Gonsenhauser says it’s encouraging that people are willing to continue these practices and that this year’s flu season is proof of their effectiveness.
“Flu cases and hospitalizations are way down compared to recent years. A lot of that is likely because precautions like masking, physical distancing and hand hygiene are working to prevent the flu,” Gonsenhauser said. “I think a lot of people realize what we’ve learned from COVID-19 can be applied more generally to keep our population healthy.”
After nearly a year of living in a world drastically changed by the pandemic, continuing these practices may ease the anxiety of returning to public spaces. Wearing a mask, for example, can provide a sense of control and comfort to those with lingering pandemic fears.
Experts also predict that there are some aspects of society that will never return to pre-pandemic standards, and that’s not all bad. Work from home options will likely stick around in many industries and the convenience of seeing your doctor virtually through a telehealth visit is likely to remain and even expand in the future.
Photo:
The Ohio State University Wexner Medical Center
Mike Nicholson livestreams his hip-hop fitness classes and has had limited in-person attendance for months. He’s looking forward to welcoming more people back to class. But to get there, everyone will have to keep following COVID-19 safety protocols.
Newswise — You might remember you ate cereal for breakfast but forget the color of the bowl. Or recall watching your partner put the milk away but can't remember on which shelf.
A new Northwestern Medicine study improved memory of complex, realistic events similar to these by applying transcranial magnetic stimulation (TMS) to the brain network responsible for memory. The authors then had participants watch videos of realistic activities to measure how memory works during everyday tasks. The findings prove it is possible to measure and manipulate realistic types of memory.
"On a day-to-day basis we must remember complex events that involve many elements, such as different locations, people and objects," said lead author Melissa Hebscher, a postdoctoral fellow at Northwestern University Feinberg School of Medicine. "We were able to show that memory for complex, realistic events can be improved in a safe and non-invasive way using brain stimulation."
The study was conducted on healthy young adults in a controlled laboratory setting. These methods, however, also could eventually be used to improve memory in individuals with memory disorders due to brain damage or neurological disorders, Hebscher said.
The study will be published Feb. 4 in the journal Current Biology.
A new approach to studying memory: Incorporating video
The study authors used TMS with the goal of altering brain activity and memory for realistic events. Immediately following stimulation, subjects performed a memory task while having their brains scanned using functional magnetic resonance imaging (fMRI).
Instead of showing study participants pictures or lists of words - typical practices in laboratory tests that analyze memory - participants in this study watched videos of everyday activities such as such as someone folding laundry or taking out the garbage.
"Our study used video clips that more closely replicate how memory works on a day-to-day basis," Hebscher said.
Following stimulation, study participants more accurately answered questions about the content of the video clips, such as identifying the shirt color an actor was wearing or the presence of a tree in the background.
Additionally, the study found that brain stimulation led to higher quality reinstatement of memories in the brain. Reinstatement is when the brain replays or relives an original event, Hebscher said. Following stimulation, a person's brain activity while watching a video more closely resembled their brain activity when remembering that same video.
"This is why remembering can sometimes feel like 'mental time travel,'" Hebscher said. "Our findings show that stimulation enhances this 'mental time travel' in the brain and improves memory accuracy. These findings have implications for the development of safe and effective ways to improve real-world memory."
How the study worked
The study authors used a brain imaging technique called multi-voxel pattern analysis to compare patterns of brain activity when subjects were watching a video to brain activity when subjects were remembering that same video. The scientists measured the effect of stimulation by comparing memory and brain activity following stimulation of the memory network to the same measures following stimulation of a control brain region that does not belong to the memory network.
During the memory test, subjects watched a large set of video clips and later remembered them and answered true/false questions about the content of the videos. The researchers found that memory network stimulation improved the number of questions that subjects answered correctly. It also increased reinstatement of videos in brain regions associated with visual processing.
"Follow-up studies will work to gather more reliable measures of the brain network responsible for memory in healthy subjects as well as in patients with memory disorders," Hebscher said. "Having a more reliable measurement of this network will help us more easily identify reinstatement in the brain and may help improve the effectiveness of stimulation for enhancing memory."
Newswise — Nurses play a crucial role in helping to reduce the stress experienced by family members of critically ill patients, according to an article in Critical Care Nurse (CCN).
Having a family member, regardless of their age, admitted to an intensive care unit (ICU) is a stressful event, and research has documented that such stress may contribute to depression, anxiety and posttraumatic stress disorder (PTSD).
The Society of Critical Care Medicine (SCCM) recently updated guidelines for family-centered care in ICU settings, which state that part of a nurse’s role is to assess stress among family members of a critically ill patient and to intervene to help reduce this stress.
“Nursing Interventions to Reduce Stress in Families of Critical Care Patients: An Integrative Review” responds to the SCCM guidelines by aiming to establish the state of knowledge regarding the stress experienced by families with a loved one in the ICU and to identify specific nursing interventions that may help.
Authors Valèrie Lebel, PhD, RN, and Sylvie Charette, PhD, RN, are professors in the department of nursing, at the Universitè du Quèbec en Outaouais (University of Quebec Outaouais), in Canada.
“The COVID-19 pandemic has reinforced that ICU nurses are the main point of interaction between the healthcare team and the family and that they are crucial to supporting the family through the ICU experience,” Lebel said. “A family-centered approach makes it possible to implement tailored interventions that are flexible and accommodate individual coping strategies.”
Their literature search of three databases (MEDLINE, CINAHL, and Cochrane) identified 934 research articles with select keywords related to the topic in the abstract or summary. The search covered the period from 2007, when the first SCCM family-centered care guidelines were issued, through 2019. From these, they removed duplicates and applied inclusion criteria, resulting in a total of 38 articles for the integrative review. Of these, 18 studies dealt with the neonatal ICU setting, nine were in pediatric ICUs, and 11 were in adult ICUs, most of which were medical and surgical ICUs.
The article summarizes the design and findings of all 38 research studies, with principal stressors for families and related nursing interventions. The analysis found that, in all three care settings, the sources of stress fell into four main categories:
Changes in the relationship between the patient and family
Altered appearance and behavior of the patient
Highly specialized care setting with unfamiliar medical equipment and healthcare staff
Communication and counseling with the healthcare staff
For each stressor, the researchers identified specific nursing interventions, such as using a family-centered approach to care and implementing appropriate stress-reducing interventions. Overall, their findings recommend that nursing interventions focus on valuing the role of family members in patient care, improving communication and providing accurate information.
Reducing family members’ stress during a loved one’s ICU stay may also help with efforts to reduce family post-ICU syndrome, with its mental health issues and decreased quality of life. In addition, more research is needed to develop tools to evaluate family members’ level of stress and principal stressors, as well as the most effective interventions to improve the family’s experience of intensive care.
As the American Association of Critical-Care Nurses’ (AACN’s) bimonthly clinical practice journal for acute and critical care nurses, CCN is a trusted source of information related to the bedside care of critically and acutely ill patients.
Access the article abstract and full-text PDF by visiting the CCN website at http://ccn.aacnjournals.org.