News
Newswise — Without directly invading the brain or nerves, the virus responsible for COVID-19 causes potentially damaging neurological injuries in about one in seven infected, a new study shows. These injuries range from temporary confusion due to low body-oxygen levels, to stroke and seizures in the most serious cases, say the study authors.
Led by researchers at NYU Grossman School of Medicine, the study showed no cases of brain or nerve inflammation (meningitis or encephalitis), indicating no immediate invasion of these organs by the pandemic virus, SARS-CoV-2.
While this should reassure patients, the neurological complications of COVID-19 should be taken seriously because they dramatically raise a patient’s risk of dying while still in hospital (by 38 percent), researchers say. Such adverse effects also raise a coronavirus patient’s likelihood (by 28 percent) of needing long-term or rehabilitation therapy immediately after their stay in hospital.
“The results of our study showed no signs that the coronavirus directly attacks the nervous system,” says study lead investigator Jennifer Frontera, MD. “The neurological complications seen in COVID-19 are predominately the secondary effects of being severely ill and suffering from low oxygen levels in the body for prolonged periods of time,” says Frontera, a professor in the Department of Neurology at NYU Langone Health.
Published in the journal Neurology online Oct. 5, the study closely monitored the progress of 606 COVID-19 adult patients diagnosed with brain or other nerve-related medical conditions at any of four NYU Langone hospitals in New York City and Long Island between March 10 and May 20, when coronavirus infections were at their peak in the region.
Frontera says that ahead of the pandemic, dozens of NYU Langone neurologists and trainees had deployed across its medical centers to assist with the expectant surge of COVID-19 patients.
Early reports from Asia and Europe, where infections had spiked before rising in the United States, she says, had also “raised the alarm” about possible brain damage from coronavirus infection. Because of this, the research team was ready to look for any signs of neurological dysfunction among the thousands of patients being admitted to hospital in the spring. Among all the hospitals, 4,491 patients tested positive for COVID-19 during that time.
Among the study’s other key results was that common neurological problems, such as confusion caused by chemical electrolyte imbalances, severe infection or kidney failure, usually arose within 48 hours of developing general COVID-19 symptoms, including fever, difficulty breathing, and cough.
Half of those neurologically affected were over the age of 71, which researchers say is significantly older than the other 3,885 patients with COVID-19 (at a media age of 63) who did not experience brain dysfunction. Most were men (66 percent) and white (63 percent). Frontera notes that the study results do suggest that Blacks are not at greater risk of neurological complications than other COVID-19 patients, which is “welcome news,” given that Blacks are widely known to be at greater risk of death from coronavirus infection. However, she says this potentially important observation requires further investigation.
While the coronavirus is known to attack other organs, including blood vessels and the heart, researchers say its main target is the lungs, where it makes breathing difficult, starving the body of oxygen it needs to stay alive. Low levels of oxygen in the body and brain was another common neurological problem, study results showed, that could lead to confusion, coma, or permanent brain damage.
“Our study results suggest that physicians need to be more aggressive in stabilizing body oxygen levels in patients with COVID-19 as a potentially key therapy for stopping, preventing and/or possibly reversing neurological problems,” says study senior investigator Steven Galetta, MD.
Galetta, the Philip K. Moskowitz, MD Professor and chair of the Department of Neurology at NYU Langone, says various blood-oxygen-raising therapies that could possibly work against neurological problems in patients with COVID-19 include early intubation or use of heart-lung machines, called ECMO, which mechanically “clean” the blood and “deliver” oxygen into it.
Funding support for the study was provided by National Institutes of Health grant P30 AG066512 and NYU Langone.
Besides Frontera and Galetta, other NYU Langone researchers involved in this study are Sakinah Sabadia, MD; Rebecca Lalchlan, DO; Taolin Fang, MD; Brent Flusty, DO; Patricio Millar-Vernetti, MD; Thomas Snyder, MD; Stephen Berger, MD; Dixon Yang, MD; Andre Granger, MD; Nicole Morgan, MD; Palek Patel, MD; Josef Gutman, MD; Kara Melmed, MD; Shashank Agarwal, MD; Mathew Bokhari, MD; Kaitlyn Lillemoe, MD; Daniel Friedman, MD; David Friedman, MD; Manisha Holmes, MD; Joshua Huang, MSc; Sujata Thawani, MD; Jonathan Howard, MD; Nada Abou-Fayssal, MD; Penina Krieger, MPhil; Ariane Lewis, MD: Aaron Lord, MD; Ting Zhou, MD; D. Ethan Kahn, DO; Barry Czeisler, MD; Jose Torres, MD; Shadi Yaghi, MD; Koto Ishida, MD; Erica Scher, RN, MPH; Dimitris Placatonakis, MD, PhD; Mengling Liu, PhD; Thomas Wisniewski, MD; Andrea Troxel, ScD; and Laura Balcer, MD, MSCE. Other study co-investigators are Sherry Chou, MD, MSc; and Ericka Fink, MD, at the University of Pittsburgh; Molly McNett, RN; and Shraddha Mainali, MD, at Ohio State University in Columbus; Raimund Helbok, MD, PhD, at the Medical University of Innsbruck in Austria; Courtney Robinson, MD; Jose Suarez, MD; and Wendy Ziai, MD, at Johns Hopkins University in Baltimore, Md.; Michelle Schober, MD; and Adam de Havenon, MD, at the University of Utah in Salt Lake City; and David Menon, MD, PhD, at the University of Cambridge in the United Kingdom.
Newswise — PITTSBURGH, Oct. 12, 2020 – Variations in a gene that regulates dopamine levels in the brain may influence the mobility of elderly and frail adults, according to new research from the University of Pittsburgh Graduate School of Public Health.
These results, published today in the Journal of The American Geriatrics Society, add to a growing body of evidence hinting that lower dopamine levels could contribute to the slower, often disabling walking patterns seen in some elderly populations.
“Most people think about dopamine’s role in mobility in the context of Parkinson’s disease, but not in normal aging,” said senior author Caterina Rosano, M.D., M.P.H., professor of epidemiology at Pitt Public Health. “We were curious to see if a genetic predisposition to produce more or less dopamine was related to mobility in individuals who had some level of frailty, yet did not have dementia, parkinsonism or any other neurological condition.”
While several genetic elements control dopamine signaling, Rosano and her team focused on a gene called COMT, which breaks down dopamine to control its levels within the brain. They also considered the frailty status of participants, which is a common consequence of aging marked by a decline in physiological function, poor adjustment to stressors and a susceptibility toward adverse health outcomes. The researchers suspected that frail participants could be particularly vulnerable to COMT-driven differences in dopamine levels.
Rosano and her collaborators examined this gene in more than 500 adults above the age of 65 in Pennsylvania, North Carolina, California and Maryland, excluding any participants taking dopamine-related medications or diagnosed with Parkinson’s disease. The researchers then looked for potential links between genotype, frailty and speed.
“We found that in older, frail adults, those who have a high-dopamine genotype are more likely to maintain a faster gait and may be more resilient to mobility disablement as they age,” said Rosano.
The team discovered that frail participants with a high-dopamine COMT genotype had a 10% faster walking speed compared with participants with the low-dopamine COMT genotype.
“This 10% difference may seem small, but it could make a big difference for a person walking across a busy street while negotiating traffic,” said Rosano. “This difference is even more striking when you consider just how many complex genes influence walking.”
Rosano and study co-author Nicolaas Bohnen, M.D., Ph.D., professor of neurology and radiology at the University of Michigan School of Medicine, are working with a team of scientists at Pitt to quantify what level of dopamine could give elders greater resilience to gait-slowing and mobility disablement. Their hope is that older adults with low dopamine levels could one day receive pharmacologic supplements of dopamine to help preserve their mobility.
“There are a lot of individuals living in the community who have dopamine levels toward the lower end of normal who don't have Parkinson's disease or psychiatric conditions,” said Rosano. “If we give dopamine to these people, could we make them more resilient? That's what we don't know yet.”
In the meantime, she suggests that there are actions that seniors can take today to keep moving. She recommends that elders focus on physical activities that are enjoyable and involve both the body and the brain, especially multi-sensory activities, such as dancing or walking with a loved one.
“I love to see grandparents walking around holding hands with their grandchildren because they have to look where they are going, where the child is going, keep an eye on the surroundings and pay attention to what the grandchild is saying, all at the same time,” said Rosano. “They get an amazing multi-sensory rehab, and it's fantastic.”
Additional authors on this research include first author Shannon Mance, B.S.N., R.N., Andrea Rosso, M.P.H., Ph.D., and Stephanie Studenski, M.D., M.P.H., all of Pitt; and Joshua Bis, Ph.D., of the University of Washington.
This research was supported by the National Institutes of Health (NIH) contracts HHSN268201200036C, HHSN268200800007C, HHSN268201800001C, N01HC55222, N01HC85079, N01HC85080, N01HC85081, N01HC85082, N01HC85083, N01HC85086 and NIH grants U01HL080295, R01HL087652, R01HL105756, R01HL103612, R01HL120393, U01HL130114, R01AG023629, and DK063491. Additional support was provided by UCLA Clinical Translational Science Institute grant UL1TR00181.
CREDIT: Ric Evans/Pitt
CAPTION: Caterina Rosano, M.D., M.P.H., professor of epidemiology, University of Pittsburgh Graduate School of Public Health.
Newswise — Rockville, Md. (October 6, 2020)—Researchers have used “omics” data containing genetic profiles of drugs to identify the hormone oxytocin as a possible treatment for COVID-19, the disease caused by the novel coronavirus (SARS-CoV-2). The study is published in Physiological Genomics. It was chosen as an APSselect article for October.
Increased inflammation that leads to a “cytokine storm”—in which the body attacks its own tissues—remains one of the most serious and least understood complications of COVID-19. To date, there are no medications approved by the U.S. Food and Drug Administration to treat COVID-19, which means that “repurposing existing drugs that can act on the adaptive immune response and prevent the cytokine storm in early phases of the disease is a priority,” authors of a new study wrote.
Oxytocin, a hormone produced in the brain, is involved in reproduction and childbirth. A synthetic form of oxytocin, frequently known by its brand name Pitocin, is given by an IV to some people to help labor progress and to stop bleeding after childbirth. Oxytocin also has anti-inflammatory properties, which promote an immune response. Previous research suggests the hormone protects against toxic injury and reduces levels of inflammatory substances in the lungs. Studies have also shown that cultured human cells with reduced expression of oxytocin receptors have higher levels of inflammatory proteins and oxidative stress.
The researchers of the new study used the National Institutes of Health’s Library of Integrated Network-Based Cellular Signatures database to analyze characteristics of genes that have been treated with drugs closely related to oxytocin. They found one drug in particular, carbetocin, has similar characteristics (called a signature) to genes with reduced expression of the inflammatory markers that trigger cytokine storm in people with COVID-19. Carbetocin’s signature indicates that the drug may promote the activation of T cells, which are immune cells that play an important role in immune response. Carbetocin’s signature is also similar to that of lopinavir, an antiretroviral medication already being explored as a treatment for COVID-19. All of these factors point to the promising potential of oxytocin as a targeted treatment for coronavirus-related cytokine storms.
“Understanding the mechanisms by which [oxytocin] or the [oxytocin system] can be a new immune target is crucial,” the research team wrote. However, “safety and efficacy of intravenous oxytocin in hospitalized patients with COVID-19 remains to be assessed.”
Read the full article, “Oxytocin’s anti-inflammatory and proimmune functions in COVID-19: a transcriptomic signature-based approach,” published in Physiological Genomics. It is highlighted as one of this month’s “best of the best” as part of the American Physiological Society’s APSselect program. Read all of this month’s selected research articles.
NOTE TO JOURNALISTS: To schedule an interview with a member of the research team, please contact the APS Communications Office or call 301.634.7314. Find more research highlights in our Newsroom.
Physiology is a broad area of scientific inquiry that focuses on how molecules, cells, tissues and organs function in health and disease. The American Physiological Society connects a global, multidisciplinary community of more than 10,000 biomedical scientists and educators as part of its mission to advance scientific discovery, understand life and improve health. The Society drives collaboration and spotlights scientific discoveries through its 16 scholarly journals and programming that support researchers and educators in their work.
UCLA Fielding School of Public Health faculty co-led study that found at least 10 distinct “hotspot” mutations in more than 80% of samples of the viruses’ genomes
Newswise — LOS ANGELES (Sept. 29, 2020) – Researchers have found at least 10 distinct “hotspot” mutations in more than 80% of randomly selected SAR-CoV-2 sequences from six countries, and these genome hotspots – seen as "typos" that can occur as the virus replicates during cellular division – could have a significant impact in the fight against the COVID-19 pandemic.
“These hotspots might select for more pathogenic variants,” said Christina Ramirez, UCLA Fielding School of Public Health professor of biostatistics, a co-author of the study. “Alternatively, mutations might evolve and could prove to be less pathogenic – the virus, after all, only survives when the host survives.”
The speed at which novel SARS-CoV-2 mutants are selected and dispersed around the world may also pose issues for the development of vaccines and therapeutics, according to the study in the journal Virus Research, co-authored by Ramirez and colleagues Stefanie Weber and Walter Doerfler, both of the Institute for Clinical and Molecular Virology, Friedrich-Alexander University (FAU), Erlangen, Germany.
“One of the major scientific problems confronted with by the SARS-CoV-2 pandemic lies in our limited understanding of the interactions between the viral and the human host genomes and the latter’s defense mechanisms against this pathogen,” said Doerfler, a physician and molecular geneticist. “The results of our study will provide a platform for those who take care of patients with infections.”
RESEARCH BRIEF
FINDINGS
During worldwide spreading among human populations, at least 10 distinct hotspot mutations had been selected and found in up to more than 80% of the randomly selected sequences from 6 countries. The increasing frequency of SARS-CoV-2 mutation hotspots might select for dangerous viral pathogens. Alternatively, there might be a limit to the number of mutable and selectable sites which, when exhausted, could prove disadvantageous to viral survival. The speed, at which novel SARS-CoV-2 mutants are selected and dispersed around the world, could have implications for the development of vaccines and therapeutics, according to a new study co-authored by Christina Ramirez, UCLA Fielding School of Public Health professor of biostatistics, and colleagues.
BACKGROUND
Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) was first identified in Wuhan, China late in 2019. Nine months later (Sept. 23, 2020), the virus has infected almost 32 million people around the world and caused at least 971,000 (3.07 %) fatalities in 220 countries and territories. Research on the genetics of the SARS-CoV-2 genome, its mutants, and their penetrance, can aid future defense strategies.
METHOD
By analyzing sequence data deposited between December 2019 and end of May 2020, researchers compared nucleotide sequences of 570 SARS-CoV-2 genomes from China, Europe, the United States, and India to the sequence of the Wuhan isolate.
IMPACT
It will now be important to correlate the identified hotspot mutations with the course and outcome of individual infections in humans. This demanding problem has not yet been tackled. Hopefully, the results of our study will provide a platform for those in SARS-CoV-2 research who take care of patients with SARS-CoV-2 infections. SARS-CoV-2 has the ability to mutate and, in its course of dissemination around the world, to select for distinct signal hotspot mutations depending on high rates of genome replication and complex environmental and genetic conditions in newly invaded territories. During its intercontinental journey, the exposure of SARS-CoV-2 to the 21st century’s repertoire of medical resources may have been an additional selective force. The impact of an increase in hotspot SARS-CoV-2 mutations on immunogenesis and the prospects for vaccine development might be experienced and will have to be examined in the future.
AUTHORS
Study authors are Stefanie Weber, Christina Ramirez, and Walter Doerfler
JOURNAL
The study is published as “Signal hotspot mutations in SARS-CoV-2 genomes evolve as the virus spreads and actively replicates in different parts of the world” in the November, 2020 edition of the peer-reviewed journal Virus Research,
FUNDING
This work was conducted under the auspices of the UCLA Fielding School of Public Health; initiation of this project was undertaken by Stefanie Weber and Walter Doerfler both at the Institute for Clinical and Molecular Virology at Friedrich Alexander University (FAU) in Erlangen-Nürnberg. No external funding was provided.
The UCLA Fielding School of Public Health, founded in 1961, is dedicated to enhancing the public's health by conducting innovative research, training future leaders and health professionals from diverse backgrounds, translating research into policy and practice, and serving our local communities and the communities of the nation and the world. The school has 690 students from 25 nations engaged in carrying out the vision of building healthy futures in greater Los Angeles, California, the nation and the world.
Newswise — Testing self-collected saliva samples could offer an easy and effective mass testing approach for detecting asymptomatic COVID-19.
Scientists at Hokkaido University and colleagues in Japan have demonstrated a quick and effective mass testing approach using saliva samples to detect individuals who have been infected with COVID-19 but are still not showing symptoms. Their findings were published in the journal Clinical Infectious Diseases.
"Rapid detection of asymptomatic infected individuals will be critical for preventing COVID-19 outbreaks within communities and hospitals," says Hokkaido University researcher Takanori Teshima, who led the study.
Many of the world's governments are showing reluctance to re-institute full national lockdowns as second waves of COVID-19 infections loom on the horizon. Testing and tracing systems will need to be ramped up in order to detect and isolate people who have the virus as early as possible.
Teshima and colleagues tested and compared the nasopharyngeal swabs and saliva samples of almost 2,000 people in Japan who did not have COVID-19 symptoms. Two different virus amplification tests were performed on most of the samples: the PCR test, which is now well-known and widely available around the world, and the less commonly used but faster and more portable RT-LAMP test.
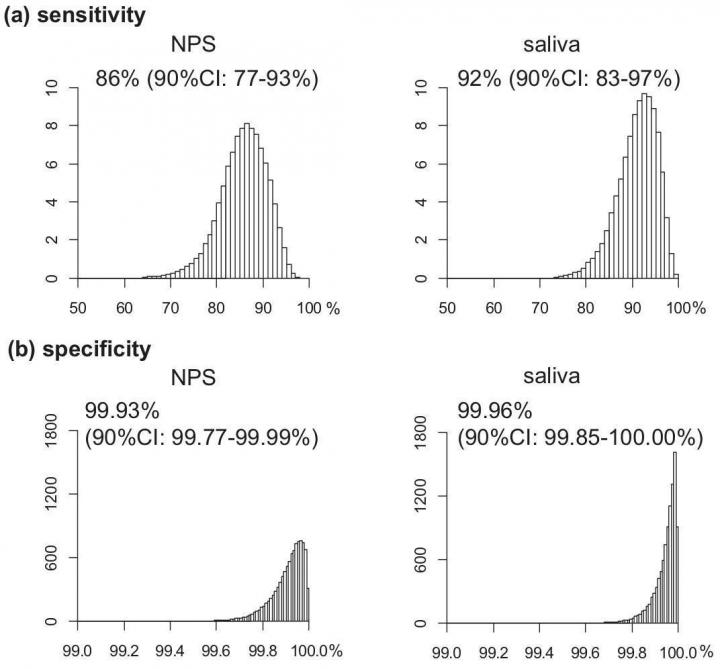
The number of positive and negative results in all samples was very similar, with the nasopharyngeal swabs and saliva samples able to detect those with the infection in 77-93% and 83-97% of subjects, respectively. Both two tests were also able to identify those without the infection in greater than 99.9% of subjects. The virus loads detected in nasopharyngeal swab and saliva were equivalent and highly correlated. Teshima says, "PCR sensitivity is much higher than previously thought 70% that came from initial data of symptomatic patients."
While finding both nasopharyngeal and saliva samples have high sensitivity and specificity to the SARS-CoV-2, Teshima says "Saliva testing has significant logistic advantages over the commonly used nasopharyngeal swab testing." "Self-collection of saliva is painless for examinees, and more importantly, it eliminates the close contact with the examiners, reducing the risk of viral exposure."
"We also found that it is unlikely that the sensitivity of RT-LAMP is significantly less than that of the PCR test, suggesting that it might be a useful alternative for diagnosing COVID-19 infection, especially where diagnosis is required at the point of sample collection, like in sports venues or at airports," says Teshima.
Researchers point to a limitation of the study that they did not follow up with clinical outcomes. Nonetheless, they suggest that the results give good indication that mass screening using self-collected saliva and rapid RT-LAMP testing could provide easy, non-invasive, quick and relatively accurate results, with minimal risk of viral transmission to healthcare workers.
Image Credit:
Isao Yokota et al., Clinical Infectious Diseases, September 25, 2020
Both nasopharyngeal swab (NPS) and saliva testing showed high sensitivity and specificity to the SARS-CoV-2. (Isao Yokota et al., Clinical Infectious Diseases, September 25, 2020)
Newswise — From Weight Watchers to wearable tech – wherever we look, there are messages encouraging us to stay fit and healthy. But diets and training methods aside, when it comes to heart health, research from the University of South Australia shows that a far more personalised approach is needed…and it all starts with your genes.
Conducted in partnership with the University of New England and the University of Queensland, the study assessed the impact of lifestyle factors on cardiovascular disease (CVD), finding clear links between genetic predisposition of CVD and smoking, alcohol intake, physical activity and diet.
UniSA researcher, Associate Professor Hong Lee, leader of the statistical genetics group at the Australian Centre for Precision Health, says the popular ‘one-size-fits-all’ approach to heart health does not have uniform effects, and that a tailored, individualised approach to CVD is essential.
Globally, CVD is the number one cause of death, claiming an estimated 17.9 million lives a year. Most deaths are due to heart attacks and strokes, with a third of these occurring prematurely in people under 70 years of age. In Australia, heart disease kills one Australian every four minutes.
“Every day, we’re exposed to information that promotes positive lifestyle factors for better health. But what we don’t hear, is how individual genetic differences can negate positive effects, often to detriment of the individual,” Assoc Prof Lee says.
“Between 20 to 60 per cent of risk factors for CVD are attributed to genetics which are far better addressed through personalised and individual interventions than broad-stroke lifestyle adjustments.
“For example, genetics show how the level of your cholesterol can be controlled by a lifestyle modification, given your genotypes and the underlying genetic link between cholesterol and lifestyle factors.
“This will help you make a decision about which lifestyle intervention is most suitable for you, for example, more exercise might be a better choice than reducing smoking.
“However, this does not necessary mean that exercise is uniformly recommended for other people who may have different genes and genetic effects that are more sensitive to smoking exposure.
“It’s all about understanding how individual genetic risks can change in line with different lifestyle adjustments, and consequently how cardiovascular health can benefit.”
Using a novel whole-genome approach, researchers analysed 23 cardiovascular health-related traits and 22 lifestyle characteristics using the ARIC (Atherosclerosis Risk in Communities) Study (N=8291) and validating results via the UK Biobank (N~500,000). 34 significant CVD trait-lifestyle pairs were identified.
While Assoc Prof Lee agrees that positive lifestyle changes are good for overall health, including cardiovascular health, he says tailored interventions based on individual differences will be most successful for managing CVD.
“As precision health practices advance, we are likely to see more personalised health treatments that are based on individual genetic profiles,” Assoc Prof Lee says.
“We are currently in the process of developing tools that can predict genetic risk based on genotypes and how lifestyle changes can modulate these.
“Incorporating individual (genetic) differences into CVD interventions will absolutely increase the predictive power of lifestyle changes on individual health.”
SAN MARCOS, TX – R-Water announces Gary Rizzato has joined the company as Chief Operating Officer (COO). In his new role, Gary will draw from experiences within the healthcare industry where he managed expansion and build out projects surpassing $1 billion and oversaw facilities totaling more than 2.5 million square feet. He will closely collaborate with CEO Rayne Guest to ensure the company's rapid growth stays aligned with the founding values and culture.
“I’ve been impressed with Gary since we met in 2015,” said Rayne Guest, founder and CEO of R-Water. “Gary’s commitment to quality care, operational efficiency, and customer service make him a perfect fit for our team.”
Gary Rizzato has over a decade of experience working in healthcare facility management and support services. Most recently, he served as Vice President for the Government Division of HHS, a company serving more than 600 partner organizations in the fields of healthcare, resorts, senior living, government, aviation, and education. Gary also assisted in starting up and managing the HHS Integrated Facilities Management service line, serving in the role of Vice President of Integrated Facilities Management.
Gary began his career working for healthcare systems, such as Tenet Health, Baton Rouge General Medical Center, and CHI St. Joseph Health. His various roles included Director of Facility Management, Property Manager, and Project Manager.
“I am proud to join R-Water, said Gary Rizzato. “The mission of the company is one I whole-heartedly believe in and look forward to using my expertise to spread the positive impact it is making in the cleaning industry even further.”
Gary received his bachelor’s degree in Environmental Design from Texas A&M University and his Master of Science in Engineering Science from Louisiana State University. He is a national design member with the American Institute of Architects (AIA), Certified Healthcare Facilities Manager (CHFM), Certified Healthcare Fire Safety Professional (CHFSP), Certified Healthcare Safety Professional (CHSP), Registered Environmental Services Executive (RESE), and possesses health and safety certifications OSHA 30 and EM 385-1-1.
About R-Water
R-Water is a woman-owned business based in San Marcos, Texas. R-Water’s computerized device gives hospitals, hotels, cruise ships, office buildings, restaurants, schools, and other facilities the power to produce cutting-edge cleaning and disinfecting solutions on-site. To learn more about how you can protect yourself against the threat of COVID-19, visit www.r-water.com or contact info@r-water.com.
Newswise — In a paper published in the Journal on Active Aging, University of Illinois Chicago longevity researcher S. Jay Olshansky and his colleagues conclude that both 2020 presidential candidates — former Vice President Joe Biden, 77, and President Donald Trump, 74 — are likely to maintain their health beyond the end of the next presidential term.
As a result, they say that chronological age and fitness should not be factors in the 2020 election.
“It is our conclusion that chronological age is not a relevant factor for either candidate running for President of the United States,” the authors write. “Both candidates face a lower than average risk of experiencing significant health or cognitive functioning challenges during the next four years.”
To evaluate each candidate’s likelihood of surviving a four-year term in office, the researchers scientifically evaluated the candidates’ health status based on publicly available medical records and confirmed publicly available personal information. The medical records of each candidate were independently evaluated by three medical doctors with experience in aging and a team of research scientists with expertise in epidemiology, public health, survival analysis, and statistics.
This is the first time that the medical records and personal attributes of presidential candidates have been scientifically evaluated by physicians and scientists in the field of aging.
The key findings of the study:
Biden and Trump are likely to be “super-agers,” a subgroup of people that maintain their mental and physical functioning and tend to live longer than the average person their age.
Both candidates have a higher than average probability of surviving a four-year term in office, relative to other men their age. For Biden, the probability of surviving the next four years is 95.2% (vs. 82.2%). For Trump, this is 90.3% (vs. 86.2%).
Biden is expected to outlive Trump, even though he is three years older. In the paper, the researchers note Biden’s “nearly perfect health profile for a man his age,” compared with Trump’s “significant but modifiable” risk factors.
While Trump is noted to have an elevated familial risk of late-onset Alzheimer’s disease, neither candidate is expected to have major cognitive functioning challenges now or during the next four years.
Olshansky, the corresponding author of the study, says the results are evidence that age does not matter in this historic election in which the next elected president will be the oldest in American history.
“We see chronological age as a topic of discussion time and again during elections, even though scientific and medical evidence tells us that biological age is far more important,” said Olshansky, professor of epidemiology and biostatics at the UIC School of Public Health.
Biological age is reflective of how rapidly a body is growing old — this occurs at different rates, Olshansky said. “Biological age is influenced by genetics and behavioral risk factors. Some people can be biologically old at age 50 while others can be biologically young at age 80.”
In prior research, Olshansky conducted the first scientific evaluation of presidential longevity; he sought to understand if being President causes an individual to age more rapidly and die sooner than expected. In that study, Olshansky concluded that most U.S. presidents actually live beyond the average life expectancy.
The new study is the first to evaluate individuals, before they are elected.
“Despite the science, the candidates themselves and their campaigns are still trying to weaponize age,” Olshansky said. “This is certainly the case for both campaigns in 2020. Comments from Biden implying that Trump is ‘mentally deranged’ and Trump’s references to Biden as ‘Sleepy Joe’ suggest that their opponents are too old, are unfit, or are otherwise unable to do the job, based on their age. It’s ageism, pure and simple.”
Tolerance of ageism, Olshansky says, harms everyone.
“We live in an aging society, and it’s important that we value, respect and continue to have a place in our culture for people of all ages. No one should be discounted from any position, even the presidency, based on their age,” Olshansky said.
Olshansky thinks that the public would be better served if age was diffused as a factor in elections rather than weaponized, and he’s seen other candidates refuse to contribute to an ageist narrative.
“Ronald Regan did this in the 80s and Pete Buttigieg did it last year. Age should not be a topic in 2020,” Olshansky said.
In 1984, Ronald Regan, then age 73, when asked about his advanced age, said “I want you to know that also I will not make age an issue of this campaign. I am not going to exploit, for political purposes, my opponent’s youth and inexperience.”
Similarly, when asked about his relative youth, Sound Bend, Indiana Mayor Pete Buttigieg, who ran in the 2019 democratic presidential primary, then age 37, deflected the question.
“Mayor Buttigieg said it’s the age of the ideas that matter, not the candidate — and I think that was right, too,” Olshansky said. “We can acknowledge age in an election, but all ages should be valued for the diverse perspectives and experience they bring.”
Co-authors on the paper are Hiram Beltrán-Sánchez of the University of California, Los Angeles; Yang Claire Yang of the University of North Carolina at Chapel Hill; Yi Li of the University of Macao; Dr. Nir Barzilai of the Albert Einstein College of Medicine; Dr. Paola Rode of Lapetus Solutions; and Dr. Bradley Willcox of the University of Hawaii.
Newswise — Middle-aged adults who report symptoms of insomnia and are sleeping less than six hours a night may be at increased risk of cognitive impairment, according to a study by Penn State College of Medicine researchers. The results may help health care professionals understand which patients who report insomnia are at increased risk for developing dementia.
Insomnia is characterized by reports of difficulty falling asleep, difficulty staying asleep or waking up too early and not being able to get back to sleep. When these symptoms occur at least 3 nights a week and for at least 3 months, it is considered a chronic disorder. Researchers found that adults who reported insomnia and obtained less than six hours of measured sleep in the laboratory were two times more likely to have cognitive impairment than people with the same insomnia complaints who got six or more hours of sleep in the lab. The study results were published in the journal SLEEP on September 24.
According to Julio Fernandez-Mendoza, associate professor of psychiatry and behavioral health and sleep specialist at Penn State Health Sleep Research and Treatment Center, about 25% of the adult general population reports insomnia symptoms and another 10% suffers from chronic insomnia. He said that being able to distinguish which of these individuals are at risk for further adverse health conditions is critical.
“This study reinforces the need to objectively measure the sleep of adults who complain of insomnia,” Fernandez-Mendoza said. In previous research, the team found that adults with insomnia who obtained less than six hours of sleep were at risk for various cardiometabolic conditions, including hypertension, diabetes, heart disease or stroke and mental health problems, such as depression.
“These new results demonstrate that these middle-aged adults also have an increased risk of cognitive impairment, which can be an early indicator of future dementia in a significant proportion of them,” Fernandez-Mendoza said.
Researchers examined data from the Penn State Adult Cohort, a randomly-selected, population-based sample of 1,741 adults who had one measured night of sleep. Before having their sleep measured in a sound, light and temperature-controlled room, participants completed a clinical history, physical exam and questionnaire to identify self-reported sleep disorders, physical health conditions, mental health problems and substance use. They also were evaluated for cognitive impairment before sleeping in the laboratory including tests that assessed attention, memory, language and other measures.
Fernandez-Mendoza and colleagues found that adults who reported insomnia symptoms or chronic insomnia and slept less than six hours in the lab were two times more likely to have cognitive impairment when compared to good sleepers. They also found that this association was particularly strong for adults with coexisting cardiometabolic conditions and cognitive impairment, which may be an indicator of vascular cognitive impairment – a condition where poor cardiovascular health results in impaired brain function.
Adults who reported insomnia but who slept six or more hours in the lab were not at risk of cognitive impartment when compared to good sleepers. The research team accounted for potential differences in sociodemographic factors – including age, sex, race, ethnicity, years of education – and the presence of physical and mental health problems, including sleep apnea, as well as substance use, such as smoking and alcohol intake.
Fernandez-Mendoza said that only having one measured night of sleep limited the study’s conclusion to in-lab sleep studies and cautioned that these data do not prove causality. Nevertheless, they further show that insomnia, cognitive impairment and cardiometabolic conditions, like high blood pressure, diabetes and heart disease, often tend to co-occur in adults who get less than six hours of sleep in the lab but not in those who can sleep six hours or more, he highlighted.
“This study is important because it is the first large U.S. prospective study associating insomnia with cognitive risk,” said Michael Twery, director of the National Center on Sleep Disorders Research of the National Heart, Lung, and Blood Institute, part of the National Institutes of Health and one of the study’s funders. “Recent scientific advances indicate that the brain depends on sleep. Understanding the connection between sleep deficiency and early cognitive decline could lead to improved treatments for insomnia.”
Fan He, Kristina Puzino, Gregory Amatrudo, Susan Calhoun, Duanping Liao, Alexandros Vgontzas and Edward Bixler of Penn State College of Medicine also contributed to this research.
The researchers declared no conflict of interest.
This research is supported in part by the American Heart Association (AHA) under award number 14SDG19830018 and the National Institutes of Health (NIH) under award numbers R01HL51931 and R01HL40916. The content is solely the responsibility of the authors and does not necessarily represent the official views of AHA or NIH.
About Penn State College of MedicineLocated on the campus of Penn State Health Milton S. Hershey Medical Center in Hershey, Pa., Penn State College of Medicine boasts a portfolio of nearly $100 million in funded research. Projects range from development of artificial organs and advanced diagnostics to groundbreaking cancer treatments and understanding the fundamental causes of disease. Enrolling its first students in 1967, the College of Medicine has more than 1,700 students and trainees in medicine, nursing, other health professions and biomedical research in both Hershey and State College, Pa.
Newswise — MINNEAPOLIS – Supplements that claim to improve mental focus and memory may contain unapproved pharmaceutical drugs and in potentially dangerous combinations and doses, according to a new study published in the September 23, 2020, online issue of Neurology® Clinical Practice, an official journal of the American Academy of Neurology. Researchers found five such drugs not approved in the United States in the supplements they examined. The supplements are sometimes called “nootropics,” “smart drugs” or “cognitive enhancers.”
“Over-the-counter cognitive supplements are popular because they promise a sharper mind, but they are not as closely regulated as pharmaceutical drugs,” said study author Pieter A. Cohen, M.D., of Harvard Medical School in Boston, Mass. “Use of these supplements poses potentially serious health risks. Not only did we detect five unapproved drugs in these products, we also detected several drugs that were not mentioned on the labels, and we found doses of unapproved drugs that were as much as four times higher than what would be considered a typical dose.”
Cohen said the supplements could be especially risky if used in combination with prescriptions drugs or instead of seeking medical advice.
Unlike pharmaceutical drugs that must be proven safe and effective for their intended use before they are marketed to consumers, the law does not require the U.S. Food and Drug Administration (FDA) to approve dietary supplements for safety or effectiveness before they reach the consumer. The FDA takes action after the products reach the market if they are mislabeled or contain unapproved products.
For the study, researchers searched the National Institutes of Health Dietary Supplement Label Database and the Natural Medicines Database for cognitive supplements that listed drugs similar to piracetam, a drug previously found in supplements but not approved in the U.S. They were looking for analogs of piracetam, drugs with a similar but slightly different chemical structure. Analogs are sometimes introduced into supplements to circumvent laws.
Researchers identified 10 supplements, eight that promised to enhance mental function, one that was marketed as “workout explosives” and another that had the words “outlast, endure, overcome” on the label.
Researchers examined the contents of each supplement using various methods and measured quantities of each drug present.
In the 10 supplements they examined, researchers detected five unapproved drugs. Two were analogs of piracetam called omberacetam and aniracetam. The others were the unapproved drugs vinpocetine, phenibut and picamilon. The FDA has issued a warning that vinpocetine should not be consumed by women of childbearing age. While all of the risks of these drugs are not known, side effects include increased and decreased blood pressure, agitation, sedation and hospitalization.
All 10 supplements contained omberacetam, which is prescribed in Russia for traumatic brain injury and mood disorders. A typical pharmaceutical dose would be 10 milligrams (mg). The doses in a recommended supplement serving size were as high as 40 mg, four times greater than in pharmaceutical dosages.
Some supplements contained more than one unapproved drug. One product combined four of the unapproved drugs.
“With as many as four unapproved drugs in individual products, and in combinations never tested in humans, people who use these cognitive enhancement supplements could be exposing themselves to potentially serious health risks,” said Cohen. “The effects of consuming untested combinations of unapproved drugs at unpredictable dosages are simply unknown and people taking these supplements should be warned.”
Researchers also found that for those products with drug quantities provided on the labels, a majority of the declared quantities were inaccurate.
“The fact that these supplements are listed in official databases does not mean the labeling is accurate or the dosage levels of ingredients in these supplements are safe,” said Cohen. “U.S. law does not permit unapproved pharmaceuticals to be introduced into dietary supplements, but the law places the burden of eliminating those products on the FDA. The FDA has issued a series of warnings to companies selling supplements with unapproved drugs, yet such drugs remain openly listed on databases as ingredients in supplements. Our study also raises concerns regarding the quality and legality of supplements listed in supplement databases.”
One limitation of the study was that it didn’t look at all unapproved drugs that are marketed in cognitive supplements.
Learn more about brain health at BrainandLife.org, home of the American Academy of Neurology’s free patient and caregiver magazine focused on the intersection of neurologic disease and brain health. Follow Brain & Life® on Facebook, Twitter and Instagram.
When posting to social media channels about this research, we encourage you to use the hashtags #Neurology and #AANscience.
The American Academy of Neurology is the world’s largest association of neurologists and neuroscience professionals, with over 36,000 members. The AAN is dedicated to promoting the highest quality patient-centered neurologic care. A neurologist is a doctor with specialized training in diagnosing, treating and managing disorders of the brain and nervous system such as Alzheimer’s disease, stroke, migraine, multiple sclerosis, concussion, Parkinson’s disease and epilepsy.
For more information about the American Academy of Neurology, visit AAN.com or find us on Facebook, Twitter, Instagram, LinkedIn and YouTube.