News
COMING HOME RETREAT FOR VETERANS – MAY 19
(Barrington, IL) The “Coming Home” Veterans Retreat offers a reflection on the transitions to and from military life. On May 19th from 8:00 a.m. to 5:30 p.m. at Bellarmine Jesuit Retreats, 420 W. County Line Road, Barrington, veterans and their family members will spend the day with others who understand and may share their experiences. This retreat is offered at no charge for all interested individuals, regardless of their current faith practices.
Bellarmine retreat leaders understand that no two veterans and no two military families are the same. Some have seen active combat, others haven’t. Some have experienced tremendous losses. Some come back healthy and strong. Some transition back to civilian life rather easily, while others face a variety of challenges. Yet serving in the military or having a loved one wear the uniform creates a common bond.
St. Ignatius of Loyola’s spirituality is the foundation of this retreat experience. Ignatius was born into a military family in 16th century Spain. His brothers were sea captains to the Americas. Ignatius was a respected commander in the Spanish army until a cannonball broke both of his legs. While recovering, he experienced a profound spiritual conversion. He no longer sought to serve an earthly king, but the Eternal King. He no longer sought victory over earthly enemies, but to conquer the spiritual battles that raged within him.
Ignatius became an expert at helping people find meaning and direction in their lives. He believed that God is hidden in the stories of our lives. Veterans and their families are invited to gather with other men and women to share experiences, to tell whatever part of your story you’re comfortable naming. Come look for God in the place you might least expect…in the pages of your life story.
In addition to personal interactions, Major General James H. Mukoyama Jr., a nationally recognized expert, will speak on Unmasking the Truth about PTSD versus Moral Injury. Fr. Ed Max McKenzie of St. Peter’s Coop Chicago and Fr. J. Michael Sparough of Bellarmine will guide conversation and engagement. Teresa Larson of Bellarmine is a veteran spouse and will reflect on family life.
Bellarmine has two libraries, a resource center, two chapels, dining room and many places to relax. Located on 80 acres populated with woodlands and prairie grasses, outdoor
spaces include walking trails, prayer gardens, a gazebo, stations of the cross as well as many patios, chairs and benches. The Retreat is designed to give the peace and serenity needed to be still and rest in God’s loving embrace.
Endorsed by the leadership of the William Chandler Peterson American Legion Post 171 in Crystal Lake, Bellarmine retreats have been experienced by several Legion members. A recent retreatant said, "It has been over 50 years since I left Vietnam. I wish this program was available when I returned home." Legion Sr. Vice Commander Bob Dorn personally attended the retreat and said, “Connect with others and learn how to better understand yourself during this inspirational day.”
To join the retreat on May 19, visit the website and make reservations or call: Bellarmine Jesuit Retreats https://jesuitretreat.org/pages/veterans-coming-home-retreats
847-381-1261
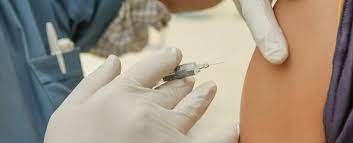
Newswise — COLUMBUS, Ohio – A simple addition to injected COVID-19 vaccines could enhance their effectiveness and provide “border protection” immunity in areas like the nose and mouth to supplement antibodies in the bloodstream, new research suggests.
The strategy involves dampening the activity of an enzyme produced by some white blood cells when they’re responding to the vaccine challenge. When highly active, this enzyme breaks down not just the pathogen – its job – but also degrades pieces of cells that participate in the immune response.
Research in mice showed that an experimental COVID-19 vaccine containing a compound to inhibit the enzyme stimulated a robust antibody response that included immunity in the nose and mouth, ultimately providing extra protection for airways and the gastrointestinal tract.
“Our approach is to improve ‘border control.’ The benefits are broad because in addition to providing protection in the bloodstream like most vaccines do, we also have excellent protection in the doors and windows of the body that communicate with the outside,” said senior study author Prosper Boyaka, professor and chair of the Department of Veterinary Biosciences at The Ohio State University.
“If we protect the mucosal area where the pathogen enters, then even if you don’t reach total immunity there, you limit the amount of pathogen that enters the body so the antibodies inside are more efficient at clearing the infection.”
The experimental vaccine was produced by packaging a segment of the SARS-CoV-2 (the virus that causes COVID-19) spike protein as an antigen with the common vaccine ingredient aluminum salts and an enzyme inhibitor. The findings suggest this affordable design could be particularly helpful in developing countries, where cold storage needed for existing vaccines is a challenge, said Boyaka, also an investigator and program director in Ohio State’s Infectious Diseases Institute.
The study was published online Aug. 5 in Proceedings of the National Academy of Sciences.
There is an irony to the use of aluminum salts (also known as alum) in about 70% of the world’s vaccines: While alum’s presence actually enhances the immune response, it also recruits the white blood cells that secrete the enzyme, called elastase.
Alum is inexpensive to obtain or produce and can be stored at room temperature, and is effective at promoting development of a bloodstream-based antibody response to vaccination. But it doesn’t do much for cell-mediated immunity that improves protection against viruses and bacteria that use cells to reproduce, and can’t generate a useful number of antibodies in the body’s portals of entry for most pathogens: the nose, mouth and genitourinary tract.
The researchers found that suppressing elastase in a vaccine containing alum had the dual benefits of broadening and speeding up the antibody response in the bloodstream and triggering the specific types of antibodies needed for immune protection of mucous membranes.
“We found a way to have the cells come and help the immune response to develop and the enzyme to break down the pathogen, but we don’t want that response to be so high that it goes out of control. So we’re just putting a brake on the activity those enzymes would have,” Boyaka said. “And we found if you apply that strategy, you can induce a response in the airways even if the vaccine is not given through the airway.”
The experimental vaccine enhanced the magnitude of mouse antibodies, which reacted to the same section of the spike protein in the vaccine that antibodies in plasma from COVID-19 patients attach to, as well as generating antibodies in mucosal areas. Immunized mice lacking the gene for the enzyme developed high-affinity antibodies as well.
To further test the concept, the researchers found the enzyme-suppressing compound used in the study triggered production of specialized inflammation-regulating cells in cultures of human immune cells and pig spleen cells, showing that this strategy could improve vaccine immune responses in other species – including people.
Boyaka’s team envisions that a future injected vaccine containing an elastase inhibitor could expand SARS-CoV-2 vaccination availability across the world and even be used to boost existing vaccines.
“COVID will stay with us for some time, unfortunately, with the new variants,” he said. “What we need to do is have a portfolio of options that we could use depending on the health environment.
“Reprogramming the immune response induced by an injected vaccine containing alum is a way to make the vaccine more efficient for what we need. This could be a cheap and simple approach that can benefit people in developing countries.”
This work was supported by grants from the National Institutes of Health and an Ohio State Office of Research COVID-19 seed grant. A patent application has been filed spanning this research; the overall patent portfolio includes an additional U.S. issued patent.
Co-authors, all from Ohio State, include Eunsoo Kim, Zayed Attia, Rachel Woodfint, Cong Zeng, Sun Hee Kim, Haley Steiner, Rajni Kant Shukla, Namal Liyanage, Shristi Ghimire, Jianrong Li, Gourapura Renukaradhya, Abhay Satoskar, Amal Amer, Shan-Lu Liu and Estelle Cormet-Boyaka.
Newswise — GRAND RAPIDS, Mich. (August 3, 2021) — The average American eats roughly 22 teaspoons of added sugar a day — more than three times the recommended amount for women and more than double the recommended amount for men.
Although this overconsumption is known to contribute to Type 2 diabetes and other disorders, the exact ways in which eating too much sugar sets the stage for metabolic diseases on a cellular level has been less clear.
Now, a team led by Van Andel Institute scientists has found that surplus sugar may cause our cellular powerplants — called mitochondria— to become less efficient, reducing their energy output.
The findings, published today in Cell Reports, highlight the cellular implications of excessive sugar consumption and provide an important new model to study the initial metabolic events that may contribute to diabetes development.
“The body needs sugar, or glucose, to survive, but, as the saying goes: ‘All good things in moderation,’” said Ning Wu, Ph.D., an assistant professor at Van Andel Institute and corresponding author of the study. “We found that too much glucose in cells, which is directly linked to the amount of sugar consumed in one’s diet, affects lipid composition throughout the body, which in turn affects the integrity of mitochondria. The overall effect is a loss of optimal function.”
Using their new model, Wu and her colleagues demonstrated that excess glucose reduces the concentration of polyunsaturated fatty acids (PUFAs) in the mitochondrial membrane and makes mitochondria less efficient. PUFAs are vital players in supporting mitochondrial function and mediating a host of other biological processes such as inflammation, blood pressure and cellular communication.
Instead, excess glucose is synthesized into a different form of fatty acid that isn’t as efficient or as flexible as PUFAs. This upends the lipid composition of the membrane and puts stress on the mitochondria, damaging them and impacting their performance.
Wu and her colleagues were able to reverse this detrimental effect by feeding their mouse models a low-sugar ketogenic diet, which suggests that reducing glucose and restoring normal membrane lipid composition supports healthy mitochondrial integrity and function. They also found that consuming excess carbohydrates reduces the beneficial effect of PUFA supplements.
“Although we may not always notice the difference in mitochondrial performance right away, our bodies do,” Wu explained. “If the lipid balance is thrown off for long enough, we may begin to feel subtle changes, such as tiring more quickly. While our study does not offer medical recommendations, it does illuminate the early stages of metabolic disease and provides insights that may shape future prevention and therapeutic efforts.”
Other authors include Althea N. Waldhart, Brejnev Muhire, Ph.D., Ben Johnson, Ph.D., Dean Pettinga, Zachary B. Madaj, M.S., Emily Wolfrum, MPH, Vanessa Wegert, and J. Andrew Pospisilik, Ph.D., of VAI; and Xianlin Han, Ph.D., of the Sam and Ann Barshop Institute for Longevity and Aging Studies and the Department of Medicine at UT Health San Antonio.
Research reported in this publication was supported by Van Andel Institute; the National Institute of General Medical Sciences of the National Institutes of Health under award no. R01GM120129 (Wu); and the National Institute on Aging of the National Institutes of Health under award no. RF1AH061872 (Han). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Newswise — WINSTON-SALEM, NC, JULY 19, 2021 -- Wake Forest researchers and clinicians are using patient-specific tumor ‘organoid’ models as a preclinical companion platform to better evaluate immunotherapy treatment for appendiceal cancer, one of the rarest cancers affecting only 1 in 100,000 people. Immunotherapies, also known as biologic therapies, activate the body’s own immune system to control, and eliminate cancer.
Appendiceal cancer is historically resistant to systemic chemotherapy, and the effect of immunotherapy is essentially unknown because clinical trials are difficult to perform due to lack of adequate patient numbers, resulting in a lack of data and limited research models.
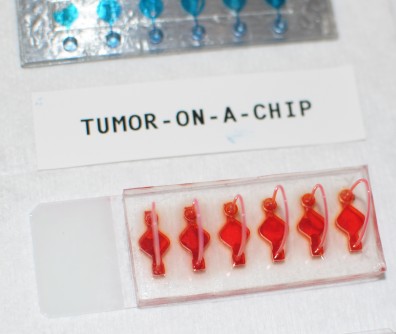
Researchers at the Wake Forest Organoid Research Center (WFORCE), a joint venture between the Wake Forest Institute for Regenerative Medicine (WFIRM), and the Wake Forest Comprehensive Cancer Center, were the first to create appendiceal cancer organoids to use as a predictive model for potential treatment options (published 2018). The Comprehensive Cancer Center is a major high volume center with a global reputation in the treatment of appendiceal cancer.
These cancer organoids are part of WFIRM’s “Body-on-a-Chip” system that allows scientists to engineer the organoids, or human tissue equivalents, that function in a very similar manner as actual human tissues and organs.
In this new study, published in the journal Clinical Cancer Research, their results indicate that various types immunotherapies tested on the organoids can potentially support treatment decisions and can achieve personalized results, identifying beneficial treatments while sparing patients from harmful side effects of drugs for which they will obtain no benefit.
“For this study we reconstructed patients’ tumors as organoids, supercharged with a built-in immune system directly obtained from the patient,” said senior author Konstantinos I. Votanopoulos, MD, PhD, professor of surgery at the Comprehensive Cancer Center and co-director of WFORCE. “In this way we created a personalized interface to study how effective the immunotherapy drugs are in activating a patient’s own immune system to kill the cancer. This platform is breaking new ground for appendiceal cancer, and it can also be applied in research for other rare cancers where preclinical models are lacking.”
This research study utilizes the WFIRM’s “Body-on-a-Chip” system that allows scientists to engineer the organoids, or humanoid tissue equivalents, that function in a very similar manner as actual human organs.
Cells from tumor biopsies from 26 patients were obtained to grow the organoids - tiny, 3D tissue-like structures, in the lab that that mimic the cancerous tumors. The immune enhanced tumor organoids were treated with one of three immunotherapy drugs and then assessed for responsiveness. “In the future, by verifying that that the tumor and its organoids behave in the same fashion, we could modify clinical trial design and optimize cost by targeting patients with organoids that have exhibited favorable results,” Votanopoulos said.
Current strategies to understand tumor progression center on analyses of the tumor cells in isolation, but do not capture the interactions between a tumor and its surrounding space, known as the microenvironment or stroma. This leads to inaccuracies in predicting tumor progression and chemotherapy or immunotherapy response. Patient-derived tumor organoids are used as a testing and predicting platform to model diseases, evaluate efficacy and/or toxicity of new and existing drugs, and can be used to test environmental hazards.
Co-author Shay Soker, PhD, professor of regenerative medicine who leads tumor organoid research at WFIRM and co-directs WFORCE, said new technologies and biological models that improve prognostication will have a significant effect on patient mortality. “Using the organoids as a preclinical platform can lead to development of novel therapeutics which target and control tumor cells specifically, sparing healthy tissue from the side effects of chemotherapy and immunotherapy treatments,” he said. “For rare cancers like appendiceal cancer, this technology can make a difference in overall quality of life for patients.”
Additional co-authors include: Steven D Forsythe, MS, Richard A Erali, MD, Shyama Sasikumar, MS, Preston Laney, BS, Ethan Shelkey, BS, all of WFIRM; and Ralph D’Agostino Jr, PhD, Lance D Miller, PhD, Perry Shen, MD, and Edward A Levine, MD, all of the Comprehensive Cancer Center. The author’s report no conflicts of interest.
Research support provided by: the Wake Forest Dean’s Hero Award, the Appendix Cancer Pseudomyxoma Peritonei Foundation, the National Organization of Rare Diseases, and the National Institute of Health. Results were first presented at the Society of Surgical Oncology Annual Meeting, March 2021.
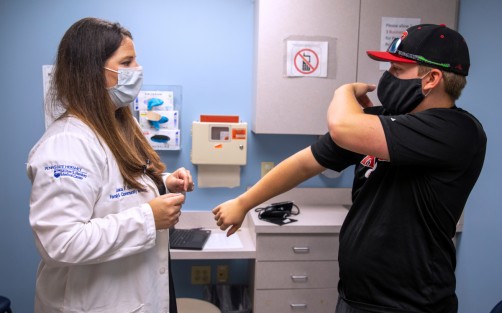
Newswise — When visiting a health care provider, most people expect to have their body temperature, pulse, weight and blood pressure measured. Some Penn State Health patients can also anticipate questions about how much they exercise. The health system is one of six in the United States and the only one in Pennsylvania to incorporate physical activity as a vital sign.
Exercise has been shown to reduce or prevent chronic illness from diabetes, high blood pressure, cardiovascular disease, certain types of cancer, stroke, dementia, depression and anxiety. Recent research also shows that it may reduce a patient’s symptom severity and risk of dying from COVID-19.
The Physical Activity Vital Sign (PAVS) involves providers asking patients questions about exercise during their pre-appointment assessment. Following screening questions about smoking or depression, patients will give information on how many minutes per day they exercise and whether that activity is light, moderate or vigorous. The answers are kept in each patient’s electronic medical record. PAVS is currently being piloted at two practices: Penn State Health Medical Group — Palmyra and Penn State Health Medical Group — Middletown. Once pilot testing is complete, Butts hopes to expand the use of PAVS to more practices across the health system.
“Many Americans don’t get enough exercise and not enough health care providers ask about or encourage it with their patients,” said Dr. Jessica Butts, assistant professor of family and community medicine and of orthopedics and rehabilitation. “By incorporating physical activity as a vital sign, it opens the door for patients and providers to discuss and incorporate exercise into treatment and prevention plans more readily.”
The U.S. Department of Health and Human Services recommends 150 minutes of moderate intensity exercise per week, 75 minutes of vigorous intensity per week or a combination of both.
Health care providers then review the physical activity responses along with other vital signs and can begin a conversation with patients about how they might include exercise as part of their treatment and prevention plans for chronic disease. In addition to improving health outcomes, Butts said preliminary research indicates that PAVS could lead to hundreds of millions of dollars in health care savings.
“Studies show that any improvement in physical activity is better than no activity at all,” Butts said. “If we can get patients to move more than they were before, our hope is that will lead to reduced levels of chronic disease, decreased health care costs and improved life expectancy.”
Penn State Health patients also have access to more than 20 exercise-related clinical trials conducted by researchers at Penn State College of Medicine. These studies examine how exercise can be used to treat conditions from fatty liver disease, joint and muscle pain to breast, liver and prostate cancers. Butts said she hopes the physical activity vital sign data might be used to study patient health outcomes and add to the existing body of evidence showing that exercise is good for health.
“The research is clear: exercise can favorably affect ailments in almost all organ systems, but it’s often undervalued and overlooked for pharmacologic or dietary interventions,” Butts said. “At Penn State Health and Penn State College of Medicine, we’re continuing to discover how physical activity might benefit the health of our patients.”
A 27-question survey of Association of American Cancer Institutes (AACI) members, including many NCI-designated centers, identified several opportunities to improve coordination of care between main centers and their satellite locations.
Newswise — PLYMOUTH MEETING, PA [July 6, 2021] — New research in the June 2021 issue of JNCCN—Journal of the National Comprehensive Cancer Network assesses the quality of cancer care delivered through extended sites coordinated by some of the country’s largest cancer centers. The study was developed to implement strategies for disseminating discoveries and expanding access to the highest quality cancer care as part of AACI’s Network Care Initiative, established by former AACI President Stanton L. Gerson, MD, Director of the Case Comprehensive Cancer Center. Results were calculated based on responses to a mixed-methods survey answered by 69 cancer centers between September 2017 and December 2018, at which time 56 reported at least one network practice site.
Just over half indicated that network sites had full access to the main centers’ electronic medical records (EMRs), and even fewer main centers had complete access to records throughout their network sites.
“Our findings demonstrate the need to improve network site alignment, particularly in patient navigators, care paths, and clinical trial access,” said Dr. Gerson, the study’s lead researcher and interim dean of the Case Western Reserve University School of Medicine. “Most federal cancer center reviews do not assess the total population of cancer patients served by major cancer centers and their affiliated sites. These data suggest that a very sizable portion of new cancer cases are cared for by these centers and their networks. Greater cancer center/network coordination could ultimately lead to improved access to clinical trials for the underrepresented communities many of these network sites serve.”
According to the survey results, some key opportunities to improve coordination of care include:
Implementing integrated EMRs across networks;
Reviewing best clinical care practices, with more rigorous use of care paths and coordination of diagnosis and treatment planning across sites;
Greater attention and support for cancer clinical trials across network sites; and
Improved physician oversight of clinical and research expectations, hiring, review and other links with cancer center main campus sites.
“Many studies show that consistency through care plans and guidelines improves patient outcomes, clinical response, and survival. More proactive approaches, including care paths, tumor boards across networks, and recognition of the value of placing disease experts at network sites, will improve the standardization of care across sites,” Dr. Gerson added.
“Disparities in cancer care outcomes, most significantly patient survival, have been shown between NCI-designated cancer centers and community hospitals, where two-thirds of cancer patients are cared for in the U.S.” commented Lawrence N. Shulman, MD, Deputy Director for Clinical Services at the Abramson Cancer Center at the University of Pennsylvania, who was not involved in this research. “Rural cancer programs often have limited cancer physicians representing all relevant specialties and urban safety-net hospitals often have limited financial resources to support high-quality cancer programs. Partnerships between academic cancer centers and community and safety-net hospitals have the potential to improve outcomes for a broader spectrum of cancer patients in the U.S. One might consider support of these cancer programs an obligation of academic cancer centers. This study outlines some potential mechanisms of support.”
The COVID-19 pandemic, which began well after this survey closed, and the growing call for increasing diversity in clinical trials, is also driving the need to better integrate network sites as a tool for delivering quality care to underserved populations. On June 7, NCCN presented a webinar on “Utilization of Network Satellite Locations” as part of a series on COVID-19 and Cancer Center Operations. Dr. Shulman was one of the panelists, along with other members of the NCCN Best Practices Committee. That video is available at: NCCN.org/covid-19.
To read the entire study, visit JNCCN.org. Complimentary access to “Status of Cancer Care at Network Sites of the Nation’s Academic Cancer Centers” is available until September 10, 2021.
As Americans are managing life with the Coronavirus and now with the vaccine finally rolling out, and mask mandates lifting for the fully vaccinated, the travel industry is continuing to come back. Hotels and travel destinations in many parts of the country are open in anticipation of travelers ready to throw off their cabin fever and venture out.
Your plans may be for a weekend get away to a cozy bed and breakfast, or perhaps a fishing trip to catch that “big one” that won’t get away this time. Maybe it’s a camping trip full of hiking adventures with stunning vistas, or possibly you would rather walk through America’s glorious past by taking in all the amenities offered in any of hundreds of museums. How about a week’s stay at a ranch out west? Whatever your travel desires are, your options are plentiful and, more importantly, clean, safe and following all CDC guidelines for Coronavirus.
Each week we will list state by state travel options we recommend to help make your vacation choices easy.
ALASKA:
Kenai Magic Lodge – Anchorage.
MD Discovery Alaskan Charters – Ketchikan.
ARIZONA:
Stampede RV and Bed & Breakfast – Tombstone.
Hall of Flame Fire Museum – Phoenix.
ARKANSAS:
Beaver Lake Hideaway – Garfield.
Urban Peddlers Retreat – VRBO-2039106 – Bentonville.
Vista del Paradiso – VRBO-1744925 – Bella Vista.
CALIFORNIA:
Mendo Parks – Mendocino.
Catalina Island Museum – Avalon.
Orangeland RV Park – Orange.
COLORADO:
Inn at Lost Creek – Telluride.
Valhalla Resort – Estes Park.
Alpine Inn – Gunnison.
Monarch Spur RV Park & Campground – Salida.
CONNECTICUT:
Connecticut Trolley Museum – East Windsor.
FLORIDA:
Pompano Beach Rentals – Pompano Beach. Barbara O’Donnell - 954.953.4992
Ramada Inn – Temple Terrace.
Light Tackle Adventures – Odessa.
El Caribe Resort and Conference Center – Daytona Beach.
GEORGIA:
National Civil War Naval Museum – Columbus.
IDAHO:
Pend Orielle Resort – Hope.
The Roosevelt Inn – Coeur D’Alene.
ILLINOIS:
Rooster Heaven Hunt Club – Forrest.
Bear Branch – Eddyville.
Keyesport Cabins – Keyesport. 618.749.5413
McLean County Museum of History – Bloomington.
INDIANA:
Grissom Air Museum – Peru.
IOWA:
Sanford Museum & Planetarium – Cherokee.
Scenic View Ranch – Monona.
KANSAS:
Combat Air Museum – Topeka.
Covered Wagon RV Resort – Abilene.
KENTUCKY:
Singing Hills RV Resort – Cave City.
Riverside Inn Bed and Breakfast – Warsaw.
Lynnhurst Family Resort – Murray.
OH Kentucky Campground and RV Park – Berea.
LOUISIANA:
City of Opelousas Tourism – Opelousas.
Cajun Country Cottages – Breaux Bridge.
Terrell House – New Orleans.
La Quinta Inn – Slidell.
MAINE:
Old Fort Western – Augusta.
York Beach Camper Park – York Bench.
MICHIGAN:
Days Inn & Suites by Wyndham – St. Ignace.
Greenwood Acres RV – Jackson.
Serendipity Bed & Breakfast – Saugatuck.
US National Ski-Snowboard Hall of Fame – Ishpeming.
Michigan Heroes Museum – Frankenmuth.
Hotel Nichols – South Haven.
MINNESOTA:
Starck’s Tamarck – Deer River.
White Oak Inn & Suites – Deer River.
Guest House International Inn & Suites – Rochester.
Kecs Kove – Kabetagoma.
MISSISSIPPI:
Magnolia Cottage Bed & Breakfast – Natchez.
MISSOURI:
KC Karting Association – Liberty.
Best Western Branson Inn – Branson.
Driftwater Resort – Branson.
MONTANA:
Montana Military Museum – Fort Harrison.
NEVADA:
Nevada Northern Railway Museum – Ely.
NEW MEXICO:
NM Holocaust Museum – Albequerque.
NEW YORK:
Adirondak Experience – Blue Mountain Lake.
Victorian Bed & Breakfast – Staten Island.
Fernbaugh Campground & Rec Center – Corning.
NORTH CAROLINA:
The Groome Inn – Greensboro.
Pinebrook Manor Bed and Breakfast – Hendersonville.
NORTH DAKOTA:
Dakota Waters Resort – Beulah.
McQuoid Outdoors & Lodging – Minnewaukan.
OHIO:
Good Earth Cabins – Logan.
OREGON:
The Fort Dalles Museum – The Dalles.
PENNSYLVANIA:
Manayunk Chambers Guest House – Philadelphia.
Inn at White Oak – Gettysburg.
TENNESSEE:
The Olde Mill Inn Bed & Breakfast – Cumberland Gap.
Mountain Breeze Motel – Pigeon Forge.
Appleview River Resort – Sevierville.
TEXAS:
Breeze Lake/Sunset Palms Campgrounds – Brownsville.
American Undersea Warfare Museum – Galveston.
National Museum of the Pacific War – Fredericksburg.
Alamo RV Park – Alamo.
Triple Creek RV Music Resort – Woodville.
VIRGINIA:
Historic Smithfield – Blacksburg.
WASHINGTON D.C.:
National Museum of the US Navy
WEST VIRGINIA:
The Lost River Grill, Motel and Bed & Breakfast – Lost River.
WISCONSIN:
Mid-Continent Railway and Museum – North Freedom.
BRITISH VIRGIN ISLANDS:
M&M Apartment Suites & Bakery – Spanish Town.
When you are ready to travel, we are here to help! Keep checking back every week for more featured locations. Our travel hosts are eager to see you and work with you to provide safe and clean facilities.
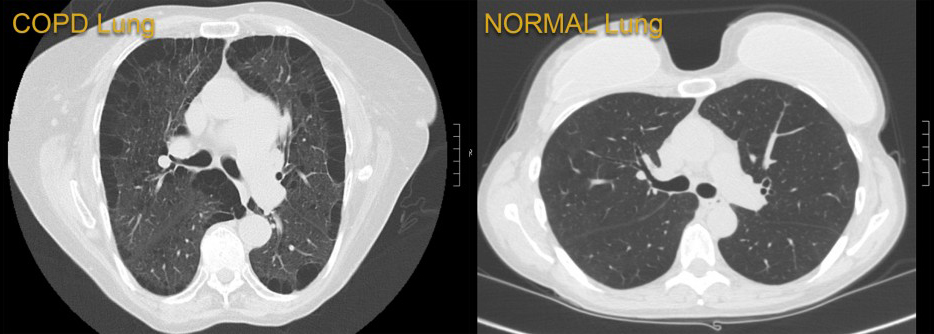
Newswise — BALTIMORE (June 14, 2021) – Researchers at the University of Maryland School of Medicine (UMSOM) analyzed data at the 13-hospital University of Maryland Medical System (UMMS) and found public health measures designed to reduce the spread of the COVID-19 virus may have fostered a substantial side benefit: Hospital admissions for chronic obstructive pulmonary disease (COPD) were reduced by 53 percent, according to a new study published in The American Journal of Medicine. This is likely due to a drop in circulating seasonal respiratory viruses such as influenza.
Hospitalizations for COPD, a group of lung diseases that make it hard to breathe and get worse over time, are commonly driven by flare-ups where symptoms are triggered by such factors as tobacco smoke, air pollution and respiratory infections. Seasonal respiratory viruses, including those that cause the common cold or influenza, trigger nearly half of those flare-ups.
In the wake of a marked drop in COPD admissions during the pandemic, the researchers theorized that COVID-19 behavior changes – a mix of stay-at-home orders, social distancing, masking mandates and strict limitations on large gatherings – not only protected against COVID-19, but they may have also reduced exposure to other respiratory infections.
Conversely, they worry that the return to normal behavior may lead to more COPD flare-ups.
“Our study shows there’s a silver lining to the behavior changes beyond protecting against COVID-19,” said senior author Robert M. Reed, MD, UMSOM Professor of Medicine and pulmonologist at the University of Maryland Medical Center (UMMC). “If we completely eliminate masks and distancing during cold and flu season, we’ll allow all those viruses that have been effectively suppressed to come raging back. There could be a lot of illness.”
Prior to the COVID-19 pandemic, COPD was the fourth-leading cause of death worldwide and a leading cause of hospital admissions in the United States. The pandemic has led to significant changes in health care delivery, including reduced admissions for COPD and other non-COVID illnesses, some of which may have stemmed from patients’ fear of contracting COVID in various hospital settings, as well as a shift toward telemedicine and outpatient COPD management during the pandemic.
To understand what may have occurred to reduce COPD admissions, the researchers compared weekly hospital admissions for COPD in the pre-COVID-19 years of 2018 and 2019, with admissions after the COVID-19 public health measures were instituted. At UMMS, those measures were implemented before April 1, 2020, so the investigators chose the same five-month period in each year for their comparison, April 1 to Sept. 30.
Co-lead author Jennifer Y. So, MD, UMSOM Assistant Professor of Medicine and COPD specialist at UMMC, said electronic medical records from multiple hospitals across a range of communities in the UMMS database facilitated a granular evaluation of changes over time. “We assessed a variety of possible causes that could affect COPD admissions including the presence of multiple diseases or medical conditions and the frequency of COPD exacerbations.”
The database findings were correlated with data on respiratory viral trends from the U.S. Centers for Disease Control and Prevention for the period of Jan. 1, 2018, through Oct. 1, 2020.
“We found a 53 percent drop in COPD admissions throughout UMMS during COVID-19. That is substantial, but equally significant, the drop in weekly COPD admissions was 36 percent lower than the declines seen in other serious medical conditions, including congestive heart failure, diabetes and heart attack,” said Dr. So.
As more and more people are vaccinated against COVID-19 and many of the public health measures of the past year are relaxed, the researchers warn that a full return to normal may again expose COPD patients to the familiar seasonal triggers.
“Our study did not assess which public health components worked to tame seasonal respiratory viruses, but a simple thing like wearing a mask while riding on public transit or working from home when you’re sick with a cold could go a long way to reduce virus exposure,” said Dr. Reed.
Dr. So, who is from South Korea, said it is a cultural norm to wear masks during the winter in her native country. “The COVID-19 pandemic has helped a lot of people around the world become more aware of the role of masking and social distancing to reduce the spread of disease,” she said.
“This is a compelling study that raises some important public health questions about protecting our most vulnerable patient populations after we are finished with the COVID-19 pandemic. I certainly think it warrants a fuller discussion,” said UMSOM Dean E. Albert Reece, MD, PhD, MBA, University Executive Vice President for Medical Affairs and the John Z. and Akiko K. Bowers Distinguished Professor.
So JY, O’Hara NN, Kenaa B, Williams JG, deBorja CL, Slejko JF, Zafari Z, Sokolow M, Zimand P, Deming M, Marx J, Pollak A, Reed RM. Decline in COPD Admissions During the COVID-19 Pandemic Associated with Lower Burden of Community Respiratory Viral Infections. The American Journal of Medicine, June 11, 2021. doi: https://doi.org/10.1016/j.amjmed.2021.05.008
Newswise — A nasal therapy, built upon on the application of a new engineered IgM antibody therapy for COVID-19, was more effective than commonly used IgG antibodies at neutralizing the COVID-19 virus in animal models, according to research recently published by The University of Texas Health Science Center at Houston (UTHealth), The University of Texas Medical Branch at Galveston (UTMB Health), the University of Houston, and IGM Biosciences, Inc.
The study was published today in Nature.
Researchers engineered IgM antibodies and found that in all cases, these antibodies were significantly more potent than standard IgG antibodies in neutralizing the COVID-19 virus. One of the engineered IgM antibodies, IGM-6268, demonstrated a significantly increased potency against the original SARS-CoV-2 and emerging variants such as the current U.K., South African, and Brazilian variants of concern (VOC) and variants of interest (VOI), as well as the antibody escape mutants for the current Emergency Use Authorization antibodies. Additionally, IGM-6268 was shown to be highly effective for prophylaxis and treatment in mouse models when administered intranasally.
“High viral load in the respiratory tract correlates with severe illness and mortality in patients with COVID-19,” said Zhiqiang An, PhD, director of UTHealth Texas Therapeutics Institute, professor and Robert A. Welch Distinguished University Chair in Chemistry at McGovern Medical School at UTHealth, and faculty member at MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences and one of the corresponding authors on the study. “Respiratory mucosal antibodies are key to clearing SARS-CoV-2 infection and reducing viral transmission and IgM antibodies are nature’s first line of defense against pathogens such as viruses.”
The current government-approved antibodies, which are all IgG antibodies, are administered intravenously at high doses and don’t directly target the main sites of viral infection.
“SARS-CoV-2 has evolved mutations that severely compromise the neutralizing activities of multiple IgG monoclonal antibodies, including those under clinical trials and authorized for emergency use. Therefore, developing new antibody therapies that can overcome these challenges is an urgent unmet need, and we are pleased with the data published today,” An said.
“Synergizing the strengths of multiple institutions from academia and industry is the key to the rapid translation from ideas to therapeutic candidates. This is another example of such success. The cross-institutional and academic-industry collaborations should be expanded to other disease indications,” said Pei-Yong Shi, PhD, professor and co-senior author of the study from the Department of Biochemistry and Molecular Biology at UTMB Health.
This antibody has been licensed to biotech partner IGM Biosciences for drug development.
“The ability to use potently neutralizing IgM antibodies against SARS-CoV-2 with broad coverage of VOCs, VOIs, and viral escape mutants, is a very exciting application of the IGM platform,” said Fred Schwarzer, CEO of IGM Biosciences. “We are grateful to our collaborators at UTHealth, UTMB Health, and our scientists at IGM for the exceptional work described in Nature today.”
Additional UTHealth authors: Zhiqiang Ku, PhD; Xiaohua Ye, PhD; Wei Xiong, MD, PhD; Junquan Liu, PhD; Ningyan Zhang, PhD; Hang Su, and Hui Deng.
Other authors include Xuping Xie, PhD; Antonio E. Muruato, PhD; Jing Zou, PhD; Yang Liu, PhD; and Vineet D. Menachery, PhD, with UTMB Health; Xinli Liu, PhD; and Sujit Biswas with the University of Houston; and Paul R. Hinton; Dean C. Ng, PhD; Yu-An Cao, PhD; Kevin B. Carlin, PhD; Elizabeth J. Haanes, PhD; Bruce A. Keyt, PhD; Stephen F. Carroll, PhD; Deepal Pandya, and Sachi Rahman with IGM Biosciences.
The work was supported by grants from the Cancer Prevention and Research Institute of Texas, the National Institutes of Health, the Welch Foundation, the Sealy Smith Foundation, the Kleberg Foundation, the John S. Dunn Foundation, the Amon G. Carter Foundation, the Gillson Longenbaugh Foundation, and the Summerfield Robert Foundation.
Photo info: Photo by UTHealth
Zhiqiang An, PhD, was one of the lead authors of a study that revealed engineered IgM antibodies were more potent than standard ones against COVID-19.
Newswise — Washington, DC (June 1, 2021) — Many individuals with kidney failure have been unable to self-isolate during the COVID-19 pandemic because they require dialysis treatments in clinics several times a week. New research that will appear in an upcoming issue of CJASN highlights the risks faced by these patients and the factors involved.
For the study, Ben Caplin, MBChB, PhD (University College London) and his colleagues, on behalf of the Pan-London COVID-19 Renal Audit Group, examined information on 5,755 patients who received dialysis in 51 clinics in London. Between March 2 and May 31, 2020, a total of 990 (17%) patients tested positive and 465 (8%) were admitted to hospitals with suspected COVID-19. COVID-19 risks were higher in patients who were older, had diabetes, lived in local communities with higher COVID-19 rates, and received dialysis at dialysis clinics that served a larger number of patients. Risks were lower in patients who received dialysis in clinics with a higher number of available side rooms and that had mask policies for asymptomatic patients. No independent association was seen with sex, ethnicity, or measures of deprivation.
“Taken together, the findings confirm the high rates of symptomatic COVID-19 among patients receiving in-center dialysis and suggest sources of transmission both within dialysis units and patients’ home communities,” said Dr. Caplin. “The work also suggests that in addition to isolation of confirmed cases, addressing factors that might reduce transmission from patients without suspected or confirmed disease might provide an additional opportunity to further modify the impact of COVID-19 in this population.”
Study co-authors include Damien Ashby, Kieran McCafferty, Richard Hull, Elham Asgari, Martin L. Ford, Nicholas Cole, Marilina Antonelou, Sarah A. Blakey, Vinay Srinivasa, Dandisonba C.B. Braide-Azikwe, Tayeba Roper, Grace Clark, Helen Cronin, Nathan J. Hayes, Bethia Manson, Alexander Sarnowski, Richard Corbett, Kate Bramham, Eirini Lioudak7, Nicola Kumar, Andrew Frankel, David Makanjuola, Claire C. Sharpe, Debasish Banerjee, and Alan D. Salama.
Disclosures: Dr. Caplin reports personal fees from Lifearc, grants from Astra Zeneca, grants from Colt Foundation, grants from Medical Research Council, from Royal Free Charity, outside the submitted work; Dr. Ashby reports personal fees from Fibrogen, outside the submitted work; Dr. Corbett has a patent WO2017148836A1: "A device for maintaining vascular connections" issued; Dr. Banerjee reports grants from AstraZeneca, grants from Kidney Research UK, personal fees from ViforPharma, outside the submitted work.
The article, titled “Risk of COVID-19 Disease, Dialysis Unit Attributes, and Infection Control Strategy among London In-Center Hemodialysis Patients,” will appear online at http://cjasn.asnjournals.org/ on June 1, 2021.
The content of this article does not reflect the views or opinions of The American Society of Nephrology (ASN). Responsibility for the information and views expressed therein lies entirely with the author(s). ASN does not offer medical advice. All content in ASN publications is for informational purposes only, and is not intended to cover all possible uses, directions, precautions, drug interactions, or adverse effects. This content should not be used during a medical emergency or for the diagnosis or treatment of any medical condition. Please consult your doctor or other qualified health care provider if you have any questions about a medical condition, or before taking any drug, changing your diet or commencing or discontinuing any course of treatment. Do not ignore or delay obtaining professional medical advice because of information accessed through ASN. Call 911 or your doctor for all medical emergencies.
Since 1966, ASN has been leading the fight to prevent, treat, and cure kidney diseases throughout the world by educating health professionals and scientists, advancing research and innovation, communicating new knowledge, and advocating for the highest quality care for patients. ASN has more than 21,000 members representing 131 countries. For more information, visit www.asn-online.org.